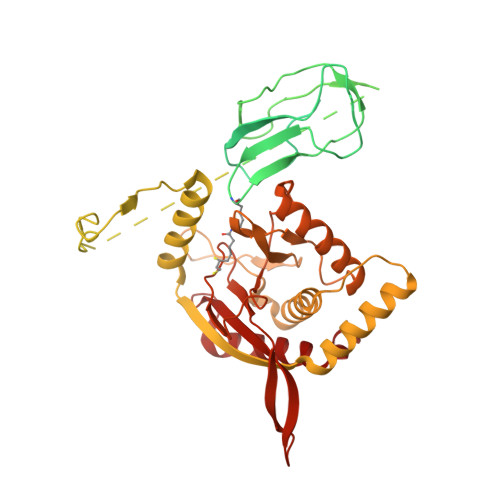
Structure of the native pyruvate dehydrogenase complex reveals the mechanism of substrate insertion.
Skerlova, J., Berndtsson, J., Nolte, H., Ott, M., Stenmark, P.(2021) Nat Commun 12: 5277-5277
- PubMed: 34489474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-25570-y
- Primary Citation Related Structures:
7B9K - PubMed Abstract:
The pyruvate dehydrogenase complex (PDHc) links glycolysis to the citric acid cycle by converting pyruvate into acetyl-coenzyme A. PDHc encompasses three enzymatically active subunits, namely pyruvate dehydrogenase, dihydrolipoyl transacetylase, and dihydrolipoyl dehydrogenase. Dihydrolipoyl transacetylase is a multidomain protein comprising a varying number of lipoyl domains, a peripheral subunit-binding domain, and a catalytic domain. It forms the structural core of the complex, provides binding sites for the other enzymes, and shuffles reaction intermediates between the active sites through covalently bound lipoyl domains. The molecular mechanism by which this shuttling occurs has remained elusive. Here, we report a cryo-EM reconstruction of the native E. coli dihydrolipoyl transacetylase core in a resting state. This structure provides molecular details of the assembly of the core and reveals how the lipoyl domains interact with the core at the active site.
- Department of Biochemistry and Biophysics, Stockholm University, Stockholm, Sweden.
Organizational Affiliation: