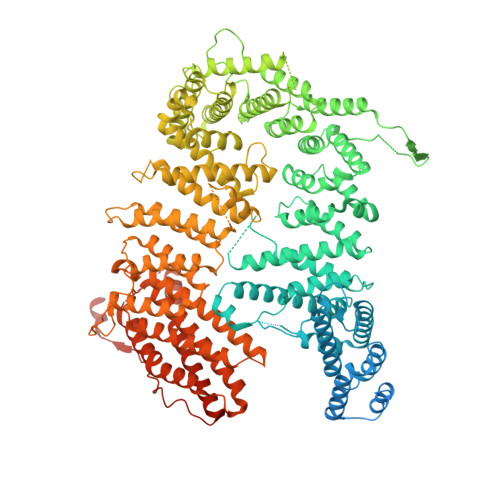
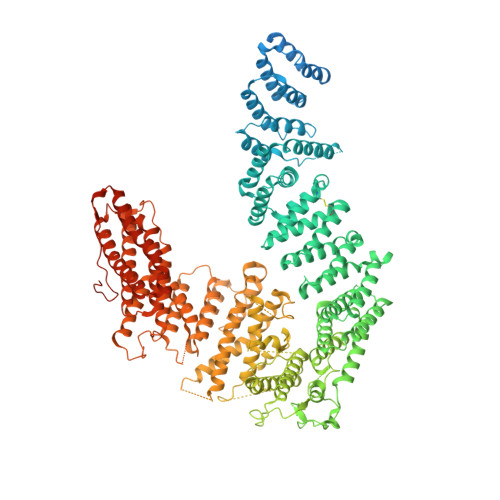
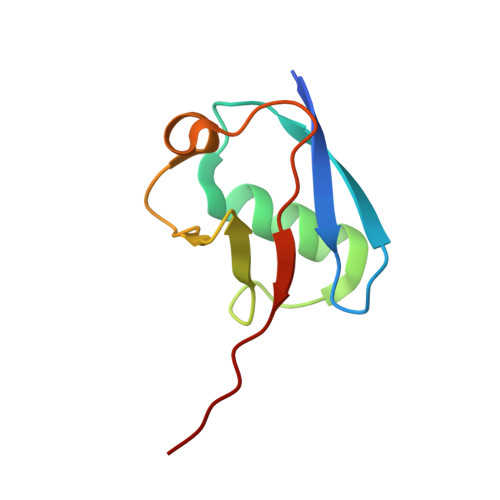
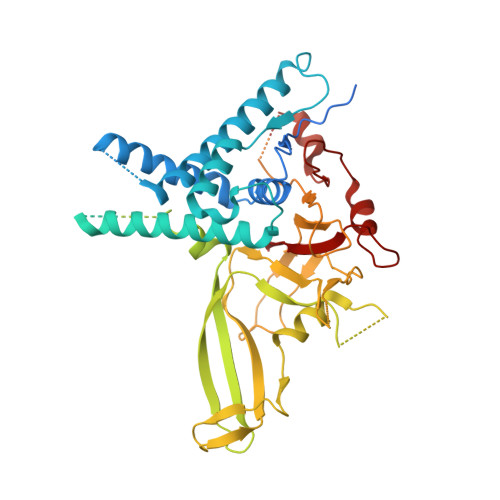
Structural basis of FANCD2 deubiquitination by USP1-UAF1.
Rennie, M.L., Arkinson, C., Chaugule, V.K., Toth, R., Walden, H.(2021) Nat Struct Mol Biol 28: 356-364
- PubMed: 33795880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00576-8
- Primary Citation Related Structures:
7AY0, 7AY1, 7AY2 - PubMed Abstract:
Ubiquitin-specific protease 1 (USP1) acts together with the cofactor UAF1 during DNA repair processes to specifically remove monoubiquitin signals. One substrate of the USP1-UAF1 complex is the monoubiquitinated FANCI-FANCD2 heterodimer, which is involved in the repair of DNA interstrand crosslinks via the Fanconi anemia pathway. Here we determine structures of human USP1-UAF1 with and without ubiquitin and bound to monoubiquitinated FANCI-FANCD2. The crystal structures of USP1-UAF1 reveal plasticity in USP1 and key differences to USP12-UAF1 and USP46-UAF1, two related proteases. A cryo-EM reconstruction of USP1-UAF1 in complex with monoubiquitinated FANCI-FANCD2 highlights a highly orchestrated deubiquitination process, with USP1-UAF1 driving conformational changes in the substrate. An extensive interface between UAF1 and FANCI, confirmed by mutagenesis and biochemical assays, provides a molecular explanation for the requirement of both proteins, despite neither being directly involved in catalysis. Overall, our data provide molecular details of USP1-UAF1 regulation and substrate recognition.
- Institute of Molecular Cell and Systems Biology, College of Medical Veterinary and Life Sciences, University of Glasgow, Glasgow, UK. Martin.Rennie@glasgow.ac.uk.
Organizational Affiliation: