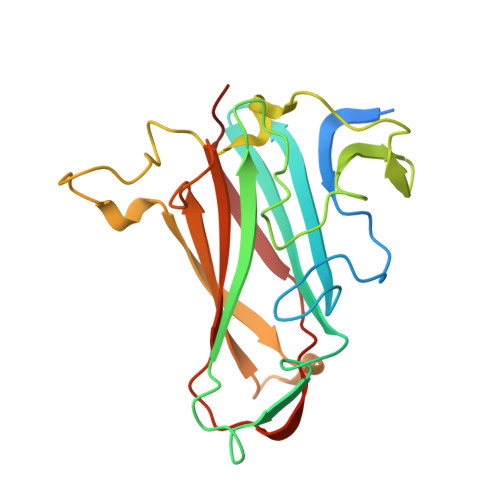
Human species D adenovirus hexon capsid protein mediates cell entry through a direct interaction with CD46.
Persson, B.D., John, L., Rafie, K., Strebl, M., Frangsmyr, L., Ballmann, M.Z., Mindler, K., Havenga, M., Lemckert, A., Stehle, T., Carlson, L.A., Arnberg, N.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33384338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2020732118
- Primary Citation Related Structures:
7AJP - PubMed Abstract:
Human adenovirus species D (HAdV-D) types are currently being explored as vaccine vectors for coronavirus disease 2019 (COVID-19) and other severe infectious diseases. The efficacy of such vector-based vaccines depends on functional interactions with receptors on host cells. Adenoviruses of different species are assumed to enter host cells mainly by interactions between the knob domain of the protruding fiber capsid protein and cellular receptors. Using a cell-based receptor-screening assay, we identified CD46 as a receptor for HAdV-D56. The function of CD46 was validated in infection experiments using cells lacking and overexpressing CD46, and by competition infection experiments using soluble CD46. Remarkably, unlike HAdV-B types that engage CD46 through interactions with the knob domain of the fiber protein, HAdV-D types infect host cells through a direct interaction between CD46 and the hexon protein. Soluble hexon proteins (but not fiber knob) inhibited HAdV-D56 infection, and surface plasmon analyses demonstrated that CD46 binds to HAdV-D hexon (but not fiber knob) proteins. Cryoelectron microscopy analysis of the HAdV-D56 virion-CD46 complex confirmed the interaction and showed that CD46 binds to the central cavity of hexon trimers. Finally, soluble CD46 inhibited infection by 16 out of 17 investigated HAdV-D types, suggesting that CD46 is an important receptor for a large group of adenoviruses. In conclusion, this study identifies a noncanonical entry mechanism used by human adenoviruses, which adds to the knowledge of adenovirus biology and can also be useful for development of adenovirus-based vaccine vectors.
- Department of Clinical Microbiology, Division of Virology, Umeå University, SE-90185 Umeå, Sweden.
Organizational Affiliation: