Selection, biophysical and structural analysis of synthetic nanobodies that effectively neutralize SARS-CoV-2.
Custodio, T.F., Das, H., Sheward, D.J., Hanke, L., Pazicky, S., Pieprzyk, J., Sorgenfrei, M., Schroer, M.A., Gruzinov, A.Y., Jeffries, C.M., Graewert, M.A., Svergun, D.I., Dobrev, N., Remans, K., Seeger, M.A., McInerney, G.M., Murrell, B., Hallberg, B.M., Low, C.(2020) Nat Commun 11: 5588-5588
- PubMed: 33149112 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19204-y
- Primary Citation Related Structures:
7A25, 7A29 - PubMed Abstract:
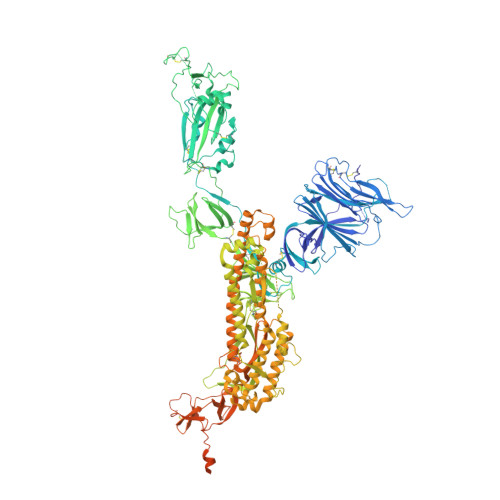
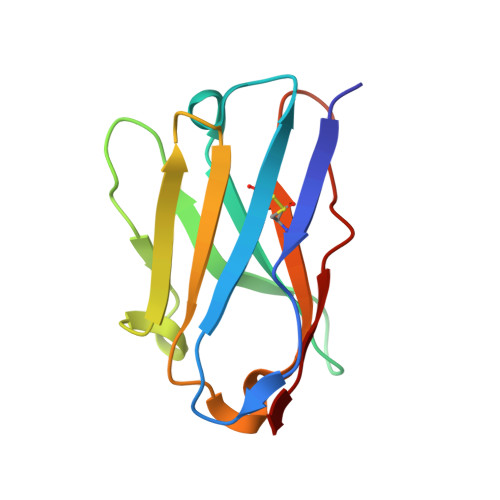
The coronavirus SARS-CoV-2 is the cause of the ongoing COVID-19 pandemic. Therapeutic neutralizing antibodies constitute a key short-to-medium term approach to tackle COVID-19. However, traditional antibody production is hampered by long development times and costly production. Here, we report the rapid isolation and characterization of nanobodies from a synthetic library, known as sybodies (Sb), that target the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein. Several binders with low nanomolar affinities and efficient neutralization activity were identified of which Sb23 displayed high affinity and neutralized pseudovirus with an IC 50 of 0.6 µg/ml. A cryo-EM structure of the spike bound to Sb23 showed that Sb23 binds competitively in the ACE2 binding site. Furthermore, the cryo-EM reconstruction revealed an unusual conformation of the spike where two RBDs are in the 'up' ACE2-binding conformation. The combined approach represents an alternative, fast workflow to select binders with neutralizing activity against newly emerging viruses.
- Centre for Structural Systems Biology (CSSB), DESY and European Molecular Biology Laboratory Hamburg, Notkestrasse 85, D-22607, Hamburg, Germany.
Organizational Affiliation: