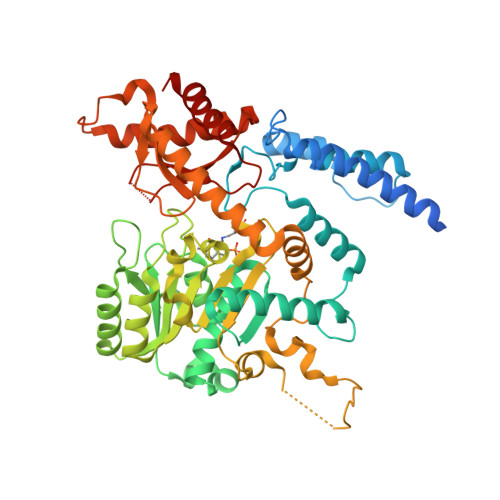
Structure and substrate specificity determinants of the taurine biosynthetic enzyme cysteine sulphinic acid decarboxylase.
Mahootchi, E., Raasakka, A., Luan, W., Muruganandam, G., Loris, R., Haavik, J., Kursula, P.(2021) J Struct Biol 213: 107674-107674
- PubMed: 33253877 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2020.107674
- Primary Citation Related Structures:
6ZEK - PubMed Abstract:
Pyridoxal 5́-phosphate (PLP) is an important cofactor for amino acid decarboxylases with many biological functions, including the synthesis of signalling molecules, such as serotonin, dopamine, histamine, γ-aminobutyric acid, and taurine. Taurine is an abundant amino acid with multiple physiological functions, including osmoregulation, pH regulation, antioxidative protection, and neuromodulation. In mammalian tissues, taurine is mainly produced by decarboxylation of cysteine sulphinic acid to hypotaurine, catalysed by the PLP-dependent cysteine sulphinic acid decarboxylase (CSAD), followed by oxidation of the product to taurine. We determined the crystal structure of mouse CSAD and compared it to other PLP-dependent decarboxylases in order to identify determinants of substrate specificity and catalytic activity. Recognition of the substrate involves distinct side chains forming the substrate-binding cavity. In addition, the backbone conformation of a buried active-site loop appears to be a critical determinant for substrate side chain binding in PLP-dependent decarboxylases. Phe94 was predicted to affect substrate specificity, and its mutation to serine altered both the catalytic properties of CSAD and its stability. Using small-angle X-ray scattering, we further showed that CSAD presents open/close motions in solution. The structure of apo-CSAD indicates that the active site gets more ordered upon internal aldimine formation. Taken together, the results highlight details of substrate recognition in PLP-dependent decarboxylases and provide starting points for structure-based inhibitor design with the aim of affecting the biosynthesis of taurine and other abundant amino acid metabolites.
- Department of Biomedicine, University of Bergen, Bergen, Norway.
Organizational Affiliation: