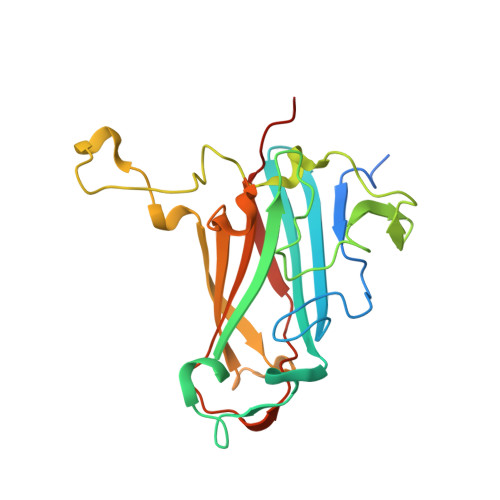
The Fiber Knob Protein of Human Adenovirus Type 49 Mediates Highly Efficient and Promiscuous Infection of Cancer Cell Lines Using a Novel Cell Entry Mechanism.
Baker, A.T., Davies, J.A., Bates, E.A., Moses, E., Mundy, R.M., Marlow, G., Cole, D.K., Bliss, C.M., Rizkallah, P.J., Parker, A.L.(2021) J Virol 95
- PubMed: 33268514 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01849-20
- Primary Citation Related Structures:
6QPO - PubMed Abstract:
The human adenovirus (HAdV) phylogenetic tree is diverse, divided across seven species and comprising over 100 individual types. Species D HAdV are rarely isolated with low rates of preexisting immunity, making them appealing for therapeutic applications. Several species D vectors have been developed as vaccines against infectious diseases, where they induce robust immunity in preclinical models and early phase clinical trials. However, many aspects of the basic virology of species D HAdV, including their basic receptor usage and means of cell entry, remain understudied. Here, we investigated HAdV-D49, which previously has been studied for vaccine and vascular gene transfer applications. We generated a pseudotyped HAdV-C5 presenting the HAdV-D49 fiber knob protein (HAdV-C5/D49K). This pseudotyped vector was efficient at infecting cells devoid of all known HAdV receptors, indicating HAdV-D49 uses an unidentified cellular receptor. Conversely, a pseudotyped vector presenting the fiber knob protein of the closely related HAdV-D30 (HAdV-C5/D30K), differing in four amino acids from HAdV-D49, failed to demonstrate the same tropism. These four amino acid changes resulted in a change in isoelectric point of the knob protein, with HAdV-D49K possessing a basic apical region compared to a more acidic region in HAdV-D30K. Structurally and biologically we demonstrate that HAdV-D49 knob protein is unable to engage CD46, while potential interaction with coxsackievirus and adenovirus receptor (CAR) is extremely limited by extension of the DG loop. HAdV-C5/49K efficiently transduced cancer cell lines of pancreatic, breast, lung, esophageal, and ovarian origin, indicating it may have potential for oncolytic virotherapy applications, especially for difficult to transduce tumor types. IMPORTANCE Adenoviruses are powerful tools experimentally and clinically. To maximize efficacy, the development of serotypes with low preexisting levels of immunity in the population is desirable. Consequently, attention has focused on those derived from species D, which have proven robust vaccine platforms. This widespread usage is despite limited knowledge in their basic biology and cellular tropism. We investigated the tropism of HAdV-D49, demonstrating that it uses a novel cell entry mechanism that bypasses all known HAdV receptors. We demonstrate, biologically, that a pseudotyped HAdV-C5/D49K vector efficiently transduces a wide range of cell lines, including those presenting no known adenovirus receptor. Structural investigation suggests that this broad tropism is the result of a highly basic electrostatic surface potential, since a homologous pseudotyped vector with a more acidic surface potential, HAdV-C5/D30K, does not display a similar pantropism. Therefore, HAdV-C5/D49K may form a powerful vector for therapeutic applications capable of infecting difficult to transduce cells.
- Division of Cancer and Genetics, School of Medicine, Cardiff University, Cardiff, United Kingdom.
Organizational Affiliation: