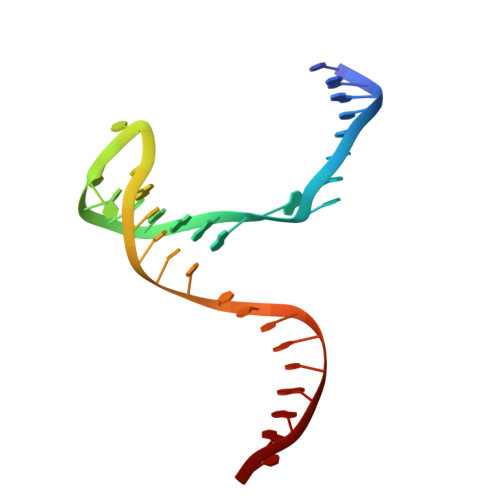
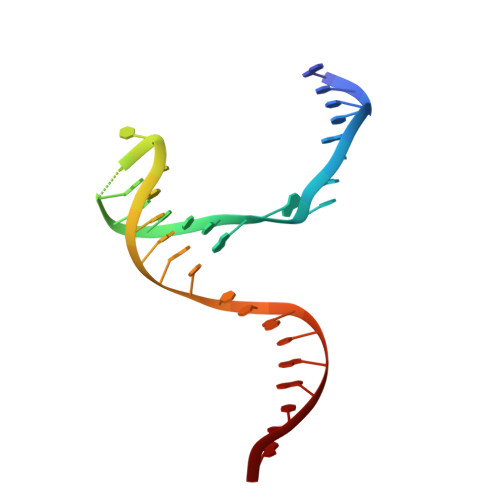
Structural basis of sequence-specific Holliday junction cleavage by MOC1.
Lin, H., Zhang, D., Zuo, K., Yuan, C., Li, J., Huang, M., Lin, Z.(2019) Nat Chem Biol 15: 1241-1248
- PubMed: 31611704
- DOI: https://doi.org/10.1038/s41589-019-0377-4
- Primary Citation Related Structures:
6IS8, 6IS9, 6JRF, 6JRG - PubMed Abstract:
The Holliday junction (HJ) is a key intermediate during homologous recombination and DNA double-strand break repair. Timely HJ resolution by resolvases is critical for maintaining genome stability. The mechanisms underlying sequence-specific substrate recognition and cleavage by resolvases remain elusive. The monokaryotic chloroplast 1 protein (MOC1) specifically cleaves four-way DNA junctions in a sequence-specific manner. Here, we report the crystal structures of MOC1 from Zea mays, alone or bound to HJ DNA. MOC1 uses a unique β-hairpin to embrace the DNA junction. A base-recognition motif specifically interacts with the junction center, inducing base flipping and pseudobase-pair formation at the strand-exchanging points. Structures of MOC1 bound to HJ and different metal ions support a two-metal ion catalysis mechanism. Further molecular dynamics simulations and biochemical analyses reveal a communication between specific substrate recognition and metal ion-dependent catalysis. Our study thus provides a mechanism for how a resolvase turns substrate specificity into catalytic efficiency.
- College of Chemistry, Fuzhou University, Fuzhou, China.
Organizational Affiliation: