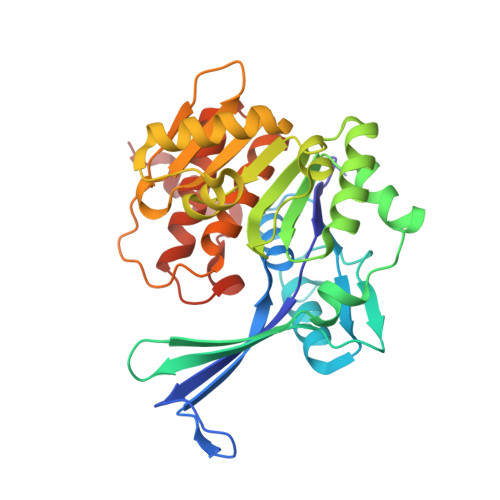
Crystal structure and mutational analyses of ribokinase from Arabidopsis thaliana.
Kang, P.A., Oh, J., Lee, H., Witte, C.P., Rhee, S.(2019) J Struct Biol 206: 110-118
- PubMed: 30822455 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.02.007
- Primary Citation Related Structures:
6ILR, 6ILS, 6ILT - PubMed Abstract:
Nitrogen remobilization is a key issue in plants. Recent studies in Arabidopsis thaliana have revealed that nucleoside catabolism supplies xanthine, a nitrogen-rich compound, to the purine ring catabolic pathway, which liberates ammonia from xanthine for reassimilation into amino acids. Similarly, pyrimidine nuclosides are degraded and the pyrimidine bases are fully catabolized. During nucleoside hydrolysis, ribose is released, and ATP-dependent ribokinase (RBSK) phosphorylates ribose to ribose-5'-phosphate to allow its entry into central metabolism recycling the sugar carbons from nucleosides. In this study, we report the crystal structure of RBSK from Arapidopsis thaliana (AtRBSK) in three different ligation states: an unliganded state, a ternary complex with ribose and ATP, and a binary complex with ATP in the presence of Mg 2+ . In the monomeric conformation, AtRBSK is highly homologous to bacterial RBSKs, including the binding sites for a monovalent cation, ribose, and ATP. Its dimeric conformation, however, does not exhibit the noticeable ligand-induced changes that were observed in bacterial orthologs. Only in the presence of Mg 2+ , ATP in the binary complex adopts a catalytically competent conformation, providing a mode of action for Mg 2+ in AtRBSK activity. The structural data combined with activity analyses of mutants allowed assignment of functional roles for the active site residues. Overall, this study provides the first structural characterization of plant RBSK, and experimentally validates a previous hypothetical model concerning the general reaction mechanism of RBSK.
- Department of Agricultural Biotechnology, Seoul National University, Seoul 151-921, Republic of Korea.
Organizational Affiliation: