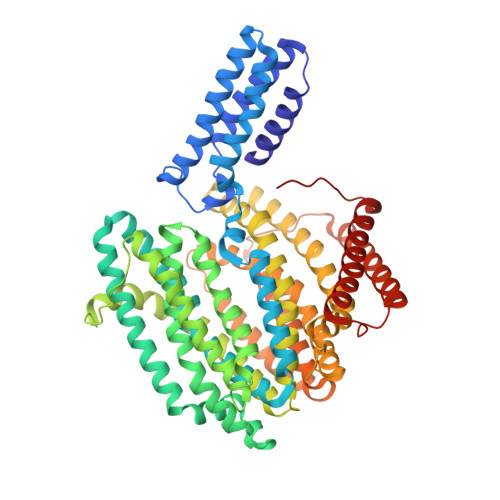
Structure and mutagenic analysis of the lipid II flippase MurJ fromEscherichia coli.
Zheng, S., Sham, L.T., Rubino, F.A., Brock, K.P., Robins, W.P., Mekalanos, J.J., Marks, D.S., Bernhardt, T.G., Kruse, A.C.(2018) Proc Natl Acad Sci U S A 115: 6709-6714
- PubMed: 29891673 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1802192115
- Primary Citation Related Structures:
6CC4 - PubMed Abstract:
The peptidoglycan cell wall provides an essential protective barrier in almost all bacteria, defining cellular morphology and conferring resistance to osmotic stress and other environmental hazards. The precursor to peptidoglycan, lipid II, is assembled on the inner leaflet of the plasma membrane. However, peptidoglycan polymerization occurs on the outer face of the plasma membrane, and lipid II must be flipped across the membrane by the MurJ protein before its use in peptidoglycan synthesis. Due to its central role in cell wall assembly, MurJ is of fundamental importance in microbial cell biology and is a prime target for novel antibiotic development. However, relatively little is known regarding the mechanisms of MurJ function, and structural data for MurJ are available only from the extremophile Thermosipho africanus Here, we report the crystal structure of substrate-free MurJ from the gram-negative model organism Escherichia coli , revealing an inward-open conformation. Taking advantage of the genetic tractability of E. coli , we performed high-throughput mutagenesis and next-generation sequencing to assess mutational tolerance at every amino acid in the protein, providing a detailed functional and structural map for the enzyme and identifying sites for inhibitor development. Lastly, through the use of sequence coevolution analysis, we identify functionally important interactions in the outward-open state of the protein, supporting a rocker-switch model for lipid II transport.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115.
Organizational Affiliation: