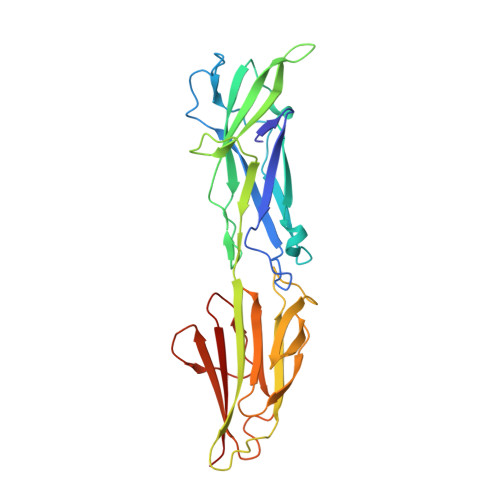
Group AStreptococcusT Antigens Have a Highly Conserved Structure Concealed under a Heterogeneous Surface That Has Implications for Vaccine Design.
Young, P.G., Raynes, J.M., Loh, J.M., Proft, T., Baker, E.N., Moreland, N.J.(2019) Infect Immun 87
- PubMed: 30936156
- DOI: https://doi.org/10.1128/IAI.00205-19
- Primary Citation Related Structures:
6BBT, 6BBW, 6N0A - PubMed Abstract:
Group A Streptococcus (GAS) ( Streptococcus pyogenes ) is an important human pathogen associated with significant global morbidity and mortality for which there is no safe and efficacious vaccine. The T antigen, a protein that polymerizes to form the backbone of the GAS pilus structure, is a potential vaccine candidate. Previous surveys of the tee gene, which encodes the T antigen, have identified 21 different tee types and subtypes such that any T antigen-based vaccine must be multivalent and carefully designed to provide broad strain coverage. In this study, the crystal structures of three two-domain T antigens (T3.2, T13, and T18.1) were determined and found to have remarkable structural similarity to the previously reported T1 antigen, despite moderate overall sequence similarity. This has enabled reliable modeling of all major two-domain T antigens to reveal that T antigen sequence variation is distributed along the full length of the protein and shields a highly conserved core. Immunoassays performed with sera from immunized animals and commercial T-typing sera identified a significant cross-reactive antibody response between T18.1, T18.2, T3.2, and T13. The existence of shared epitopes between T antigens, combined with the remarkably conserved structure and high level of surface sequence divergence, has important implications for the design of multivalent T antigen-based vaccines.
- School of Biological Sciences, The University of Auckland, Auckland, New Zealand p.young@auckland.ac.nz n.moreland@auckland.ac.nz.
Organizational Affiliation: