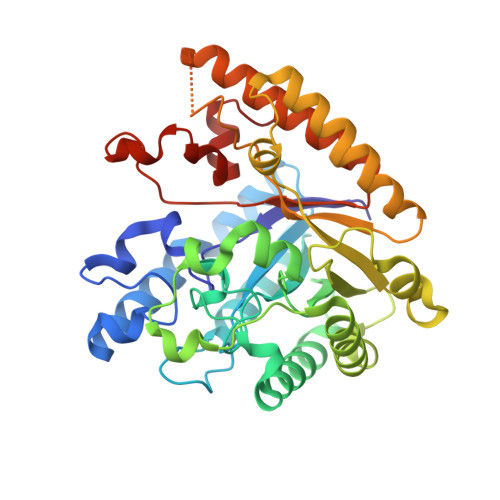
A homodimeric bacterial exo-beta-1,3-glucanase derived from moose rumen microbiome shows a structural framework similar to yeast exo-beta-1,3-glucanases.
Kalyani, D.C., Reichenbach, T., Aspeborg, H., Divne, C.(2021) Enzyme Microb Technol 143: 109723-109723
- PubMed: 33375982
- DOI: https://doi.org/10.1016/j.enzmictec.2020.109723
- Primary Citation Related Structures:
6ZB8, 6ZB9 - PubMed Abstract:
The impact of various β-glucans on the gut microbiome and immune system of vertebrates is becoming increasingly recognized. Besides the fundamental interest in understanding how β-glucans support human and animal health, enzymes that metabolize β-glucans are of interest for hemicellulose bioprocessing. Our earlier metagenomic analysis of the moose rumen microbiome identified a gene coding for a bacterial enzyme with a possible role in β-glucan metabolization. Here, we report that the enzyme, mrbExg5, has exo-β-1,3-glucanase activity on β-1,3-linked glucooligosaccharides and laminarin, but not on β-1,6- or β-1,4-glycosidic bonds. Longer oligosaccharides are good substrates, while shorter substrates are readily transglycosylated into longer products. The enzyme belongs to glycoside hydrolase subfamily GH5_44, which is a close phylogenetic neighbor of the subfamily GH5_9 exo-β-1,3-glucanases of the yeasts Saccharomyces cerevisiae and Candida albicans. The crystal structure shows that unlike the eukaryotic relatives, mrbExg5 is a functional homodimer with a binding region characterized by: (i) subsite +1 can accommodate a branched sugar on the β-1,3-glucan backbone; (ii) subsite +2 is restricted to exclude backbone substituents; and (iii) a fourth subsite (+3) formed by a unique loop. mrbExg5 is the first GH5_44 enzyme to be structurally characterized, and the first bacterial GH5 with exo-β-1,3-glucanase activity.
- Department of Industrial Biotechnology, School of Engineering Sciences in Chemistry, Biotechnology, and Health, KTH Royal Institute of Technology, SE-11421 Stockholm, Sweden.
Organizational Affiliation: