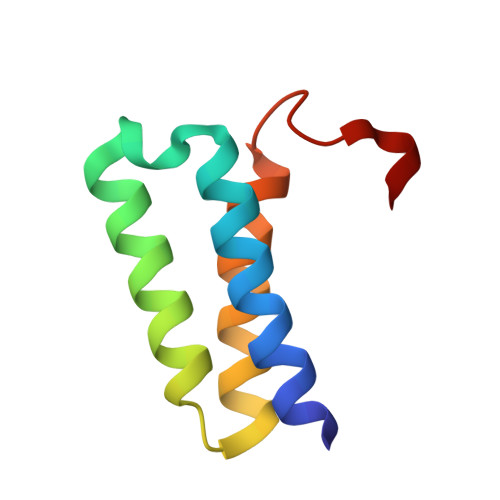
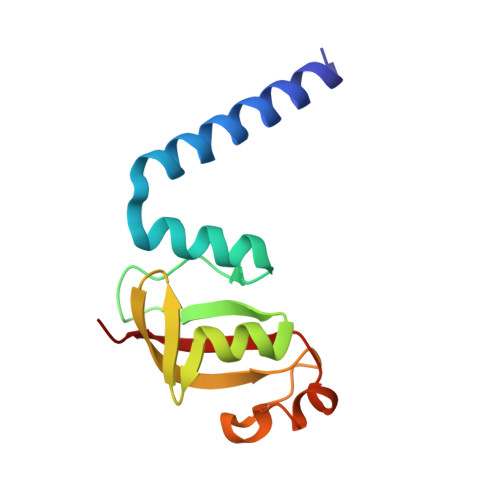
Structural analysis of the putative SARS-CoV-2 primase complex.
Konkolova, E., Klima, M., Nencka, R., Boura, E.(2020) J Struct Biol 211: 107548-107548
- PubMed: 32535228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2020.107548
- Primary Citation Related Structures:
6YHU - PubMed Abstract:
We report the crystal structure of the SARS-CoV-2 putative primase composed of the nsp7 and nsp8 proteins. We observed a dimer of dimers (2:2 nsp7-nsp8) in the crystallographic asymmetric unit. The structure revealed a fold with a helical core of the heterotetramer formed by both nsp7 and nsp8 that is flanked with two symmetry-related nsp8 β-sheet subdomains. It was also revealed that two hydrophobic interfaces one of approx. 1340 Å 2 connects the nsp7 to nsp8 and a second one of approx. 950 Å 2 connects the dimers and form the observed heterotetramer. Interestingly, analysis of the surface electrostatic potential revealed a putative RNA binding site that is formed only within the heterotetramer.
- Institute of Organic Chemistry and Biochemistry AS CR, v.v.i, Flemingovo nam. 2, 166 10 Prague 6, Czech Republic.
Organizational Affiliation: