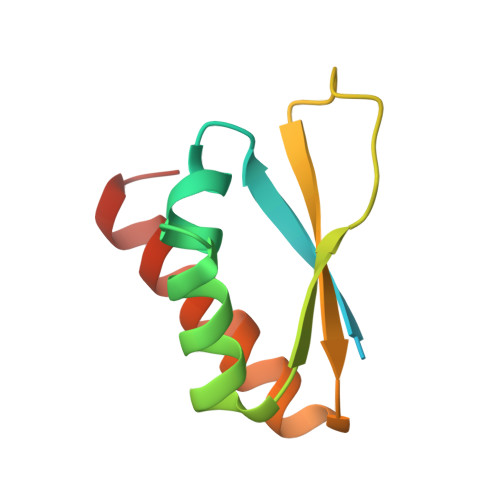
Comparative structural analyses and nucleotide-binding characterization of the four KH domains of FUBP1.
Ni, X., Knapp, S., Chaikuad, A.(2020) Sci Rep 10: 13459-13459
- PubMed: 32778776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-69832-z
- Primary Citation Related Structures:
6Y24, 6Y2C, 6Y2D - PubMed Abstract:
The FUBP1-FUSE complex is an essential component of a transcription molecular machinery that is necessary for tight regulation of expression of many key genes including c-Myc and p21. FUBP1 utilizes its four articulated KH modules, which function cooperatively, for FUSE nucleotide binding. To understand molecular mechanisms fundamental to the intermolecular interaction, we present a set of crystal structures, as well ssDNA-binding characterization of FUBP1 KH domains. All KH1-4 motifs were highly topologically conserved, and were able to interact with FUSE individually and independently. Nevertheless, differences in nucleotide binding properties among the four KH domains were evident, including higher nucleotide-binding potency for KH3 as well as diverse nucleotide sequence preferences. Variations in amino acid compositions at one side of the binding cleft responsible for nucleobase resulted in diverse shapes and electrostatic charge interaction, which might feasibly be a contributing factor for different nucleotide-binding propensities among KH1-4. Nonetheless, conservation of structure and nucleotide-binding property in all four KH motifs is essential for the cooperativity of multi KH modules present in FUBP1 towards nanomolar affinity for FUSE interaction. Comprehensive structural comparison and ssDNA binding characteristics of all four KH domains presented here provide molecular insights at a fundamental level that might be beneficial for elucidating the mechanisms of the FUBP1-FUSE interaction.
- Institute of Pharmaceutical Chemistry, Goethe-University Frankfurt, 60438, Frankfurt, Germany.
Organizational Affiliation: