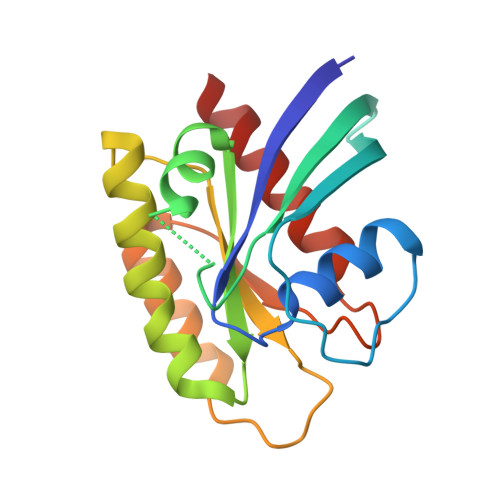
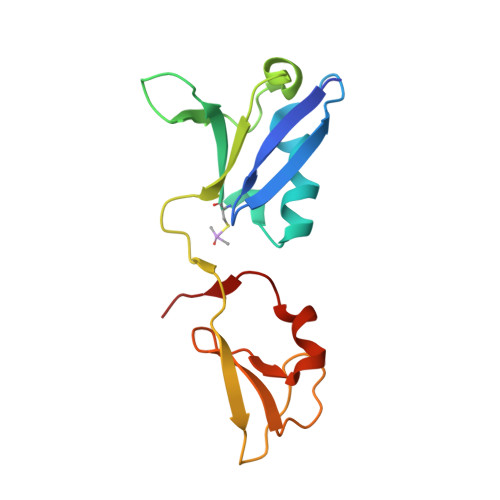
KRAS interaction with RAF1 RAS-binding domain and cysteine-rich domain provides insights into RAS-mediated RAF activation.
Tran, T.H., Chan, A.H., Young, L.C., Bindu, L., Neale, C., Messing, S., Dharmaiah, S., Taylor, T., Denson, J.P., Esposito, D., Nissley, D.V., Stephen, A.G., McCormick, F., Simanshu, D.K.(2021) Nat Commun 12: 1176-1176
- PubMed: 33608534 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-21422-x
- Primary Citation Related Structures:
6VJJ, 6XGU, 6XGV, 6XHA, 6XHB, 6XI7 - PubMed Abstract:
The first step of RAF activation involves binding to active RAS, resulting in the recruitment of RAF to the plasma membrane. To understand the molecular details of RAS-RAF interaction, we present crystal structures of wild-type and oncogenic mutants of KRAS complexed with the RAS-binding domain (RBD) and the membrane-interacting cysteine-rich domain (CRD) from the N-terminal regulatory region of RAF1. Our structures reveal that RBD and CRD interact with each other to form one structural entity in which both RBD and CRD interact extensively with KRAS. Mutations at the KRAS-CRD interface result in a significant reduction in RAF1 activation despite only a modest decrease in binding affinity. Combining our structures and published data, we provide a model of RAS-RAF complexation at the membrane, and molecular insights into RAS-RAF interaction during the process of RAS-mediated RAF activation.
- NCI RAS Initiative, Cancer Research Technology Program, Frederick National Laboratory for Cancer Research, Leidos Biomedical Research, Inc., Frederick, MD, USA.
Organizational Affiliation: