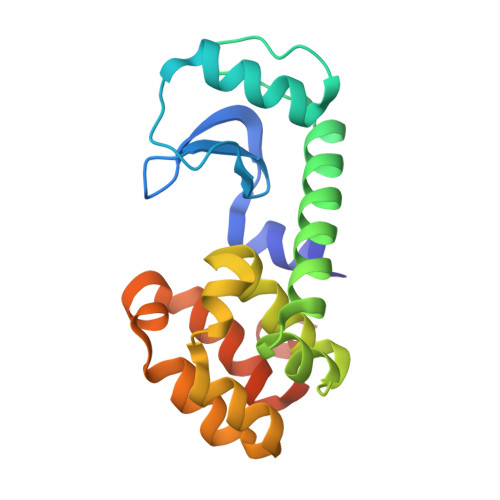
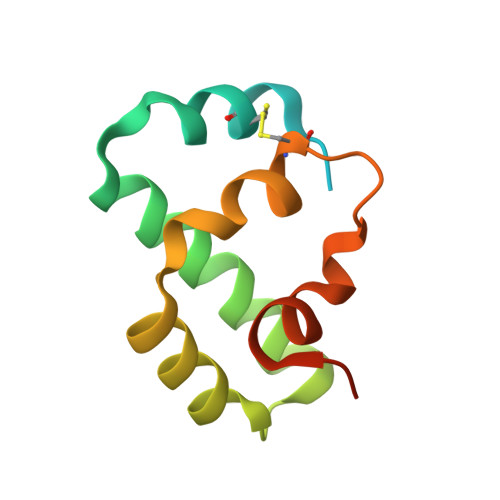
Structural basis of superinfection exclusion by bacteriophage T4 Spackle.
Shi, K., Oakland, J.T., Kurniawan, F., Moeller, N.H., Banerjee, S., Aihara, H.(2020) Commun Biol 3: 691-691
- PubMed: 33214665 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-01412-3
- Primary Citation Related Structures:
6XC0, 6XC1 - PubMed Abstract:
A bacterial cell infected with T4 phage rapidly establishes resistance against further infections by the same or closely related T-even-type bacteriophages - a phenomenon called superinfection exclusion. Here we show that one of the T4 early gene products and a periplasmic protein, Spackle, forms a stoichiometric complex with the lysozyme domain of T4 tail spike protein gp5 and potently inhibits its activity. Crystal structure of the Spackle-gp5 lysozyme complex shows that Spackle binds to a horseshoe-shaped basic patch surrounding the oligosaccharide-binding cleft and induces an allosteric conformational change of the active site. In contrast, Spackle does not appreciably inhibit the lysozyme activity of cytoplasmic T4 endolysin responsible for cell lysis to release progeny phage particles at the final step of the lytic cycle. Our work reveals a unique mode of inhibition for lysozymes, a widespread class of enzymes in biology, and provides a mechanistic understanding of the T4 bacteriophage superinfection exclusion.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, 321 Church Street S.E., Minneapolis, MN, 55455, USA.
Organizational Affiliation: