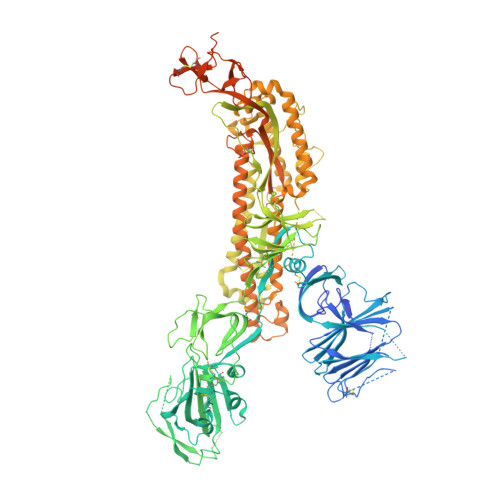
Structure-guided covalent stabilization of coronavirus spike glycoprotein trimers in the closed conformation.
McCallum, M., Walls, A.C., Bowen, J.E., Corti, D., Veesler, D.(2020) Nat Struct Mol Biol 27: 942-949
- PubMed: 32753755
- DOI: https://doi.org/10.1038/s41594-020-0483-8
- Primary Citation Related Structures:
6X79 - PubMed Abstract:
SARS-CoV-2 is the causative agent of the COVID-19 pandemic, with 10 million infections and more than 500,000 fatalities by June 2020. To initiate infection, the SARS-CoV-2 spike (S) glycoprotein promotes attachment to the host cell surface and fusion of the viral and host membranes. Prefusion SARS-CoV-2 S is the main target of neutralizing antibodies and the focus of vaccine design. However, its limited stability and conformational dynamics are limiting factors for developing countermeasures against this virus. We report here the design of a construct corresponding to the prefusion SARS-CoV-2 S ectodomain trimer, covalently stabilized by a disulfide bond in the closed conformation. Structural and antigenicity analyses show we successfully shut S in the closed state without otherwise altering its architecture. We demonstrate that this strategy is applicable to other β-coronaviruses, such as SARS-CoV and MERS-CoV, and might become an important tool for structural biology, serology, vaccine design and immunology studies.
- Department of Biochemistry, University of Washington, Seattle, WA, USA.
Organizational Affiliation: