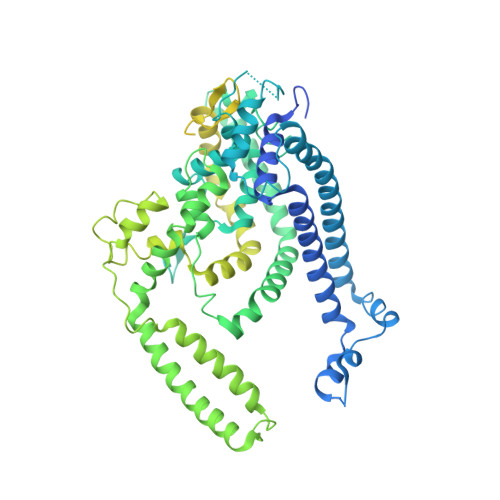
Structure of Human ATG9A, the Only Transmembrane Protein of the Core Autophagy Machinery.
Guardia, C.M., Tan, X.F., Lian, T., Rana, M.S., Zhou, W., Christenson, E.T., Lowry, A.J., Faraldo-Gomez, J.D., Bonifacino, J.S., Jiang, J., Banerjee, A.(2020) Cell Rep 31: 107837-107837
- PubMed: 32610138
- DOI: https://doi.org/10.1016/j.celrep.2020.107837
- Primary Citation of Related Structures:
6WQZ, 6WR4 - PubMed Abstract:
Autophagy is a catabolic process involving capture of cytoplasmic materials into double-membraned autophagosomes that subsequently fuse with lysosomes for degradation of the materials by lysosomal hydrolases. One of the least understood components of the autophagy machinery is the transmembrane protein ATG9. Here, we report a cryoelectron microscopy structure of the human ATG9A isoform at 2.9-Å resolution. The structure reveals a fold with a homotrimeric domain-swapped architecture, multiple membrane spans, and a network of branched cavities, consistent with ATG9A being a membrane transporter. Mutational analyses support a role for the cavities in the function of ATG9A. In addition, structure-guided molecular simulations predict that ATG9A causes membrane bending, explaining the localization of this protein to small vesicles and highly curved edges of growing autophagosomes.
- Section on Intracellular Protein Trafficking, Neurosciences and Cellular and Structural Biology Division, Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: