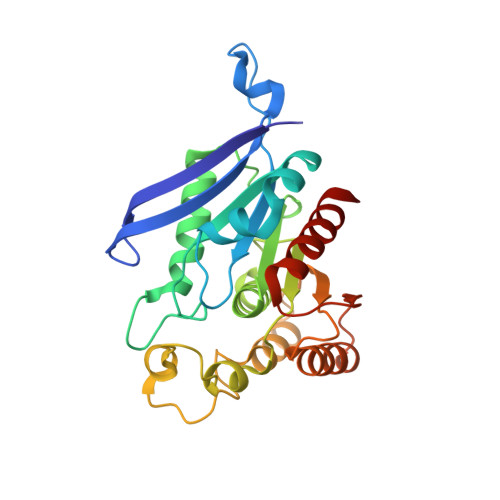
Structural Basis for the Inhibitor and Substrate Specificity of the Unique Fph Serine Hydrolases of Staphylococcus aureus .
Fellner, M., Lentz, C.S., Jamieson, S.A., Brewster, J.L., Chen, L., Bogyo, M., Mace, P.D.(2020) ACS Infect Dis 6: 2771-2782
- PubMed: 32865965 Search on PubMed
- DOI: https://doi.org/10.1021/acsinfecdis.0c00503
- Primary Citation Related Structures:
6VH9, 6VHD, 6VHE, 6WCX - PubMed Abstract:
Staphylococcus aureus is a prevalent bacterial pathogen in both community and hospital settings, and its treatment is made particularly difficult by resilience within biofilms. Within this niche, serine hydrolase enzymes play a key role in generating and maintaining the biofilm matrix. Activity-based profiling has previously identified a family of serine hydrolases, designated fluorophosphonate-binding hydrolases (Fph's), some of which contribute to the virulence of S. aureus in vivo . These 10 Fph proteins have limited annotation and have few, if any, characterized bacterial or mammalian homologues. This suggests unique hydrolase functions even within bacterial species. Here we report structures of one of the most abundant Fph family members, FphF. Our structures capture FphF alone, covalently bound to a substrate analogue and bound to small molecule inhibitors that occupy the hydrophobic substrate-binding pocket. In line with these findings, we show that FphF has promiscuous esterase activity toward hydrophobic lipid substrates. We present docking studies that characterize interactions of inhibitors and substrates within the active site environment, which can be extended to other Fph family members. Comparison of FphF to other esterases and the wider Fph protein family suggest that FphF forms a new esterase subfamily. Our data suggest that other Fph enzymes, including the virulence factor FphB, are likely to have more restricted substrate profiles than FphF. This work demonstrates a clear molecular rationale for the specificity of fluorophosphonate probes that target FphF and provides a structural template for the design of enhanced probes and inhibitors of the Fph family of serine hydrolases.
- Biochemistry Department, School of Biomedical Sciences, University of Otago, Dunedin 9054, New Zealand.
Organizational Affiliation: