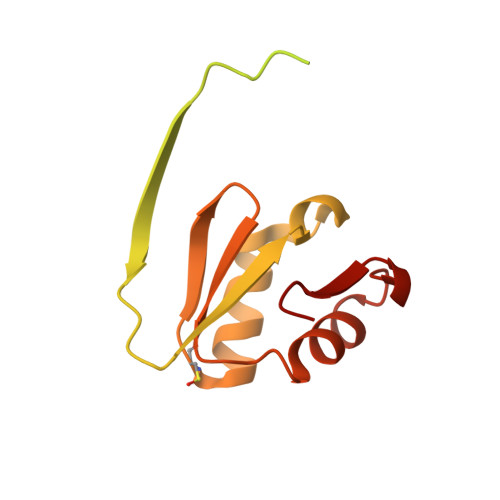
Crystal structure of the C-terminal domain of DENR.
Lomakin, I.B., De, S., Wang, J., Borkar, A.N., Steitz, T.A.(2020) Comput Struct Biotechnol J 18: 696-704
- PubMed: 32257053 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2020.03.009
- Primary Citation Related Structures:
6VPQ, 6VPR - PubMed Abstract:
The density regulated protein (DENR) forms a stable heterodimer with malignant T-cell-amplified sequence 1 (MCT-1). DENR-MCT-1 heterodimer then participates in regulation of non-canonical translation initiation and ribosomal recycling. The N-terminal domain of DENR interacts with MCT-1 and carries a classical tetrahedral zinc ion-binding site, which is crucial for the dimerization. DENR-MCT-1 binds the small (40S) ribosomal subunit through interactions between MCT-1 and helix h24 of the 18S rRNA, and through interactions between the C-terminal domain of DENR and helix h44 of the 18S rRNA. This later interaction occurs in the vicinity of the P site that is also the binding site for canonical translation initiation factor eIF1, which plays the key role in initiation codon selection and scanning. Sequence homology modeling and a low-resolution crystal structure of the DENR-MCT-1 complex with the human 40S subunit suggests that the C-terminal domain of DENR and eIF1 adopt a similar fold. Here we present the crystal structure of the C-terminal domain of DENR determined at 1.74 Å resolution, which confirms its resemblance to eIF1 and advances our understanding of the mechanism by which DENR-MCT-1 regulates non-canonical translation initiation and ribosomal recycling.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520-8114, USA.
Organizational Affiliation: