Crystal Structure of Metallo Beta Lactamase from Hirschia baltica in the Complex with the Inhibitor Captopril.
Maltseva, N., Kim, Y., Clancy, S., Endres, M., Mulligan, R., Joachimiak, A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-lactamase | 235 | Hirschia baltica ATCC 49814 | Mutation(s): 1 EC: 3.5.2.6 | 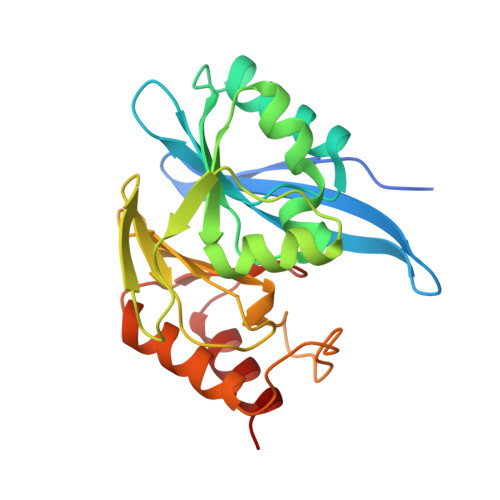 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | C6XID6 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| X8Z (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A] | L-CAPTOPRIL C9 H15 N O3 S FAKRSMQSSFJEIM-RQJHMYQMSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | B [auth A], C [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | D [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| FMT Download:Ideal Coordinates CCD File | H [auth A], I [auth A] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | J [auth A], K [auth A], L [auth A], M [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | F [auth A], G [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 77.893 | α = 90 |
| b = 77.893 | β = 90 |
| c = 241.426 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| HKL-3000 | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Human Genome Research Institute (NIH/NHGRI) | United States | -- |