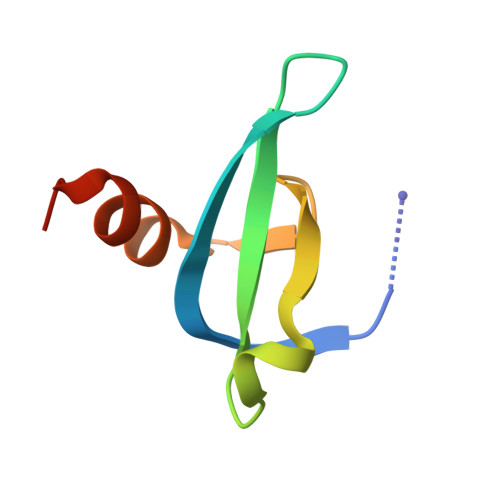
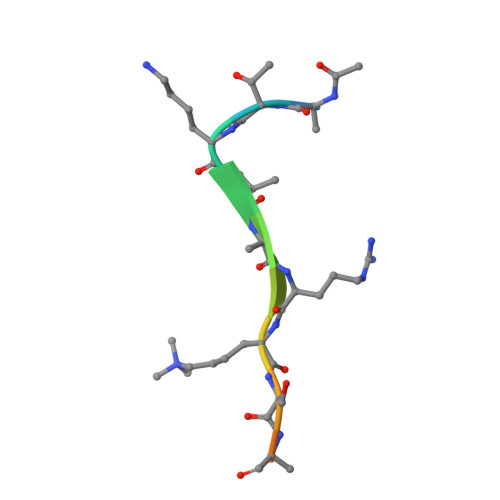
Structural Basis for the Binding Selectivity of Human CDY Chromodomains.
Dong, C., Liu, Y., Lyu, T.J., Beldar, S., Lamb, K.N., Tempel, W., Li, Y., Li, Z., James, L.I., Qin, S., Wang, Y., Min, J.(2020) Cell Chem Biol 27: 827-838.e7
- PubMed: 32470319 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2020.05.007
- Primary Citation Related Structures:
2MJ8, 6V2D, 6V2H, 6V2R, 6V2S, 6V3N, 6V41, 6V8W - PubMed Abstract:
The CDY (chromodomain on the Y) proteins play an essential role in normal spermatogenesis and brain development. Dysregulation of their expression has been linked to male infertility and various neurological diseases. Like the chromodomains of HP1 and Polycomb, the CDY chromodomains also recognize the lysine-methylated ARKS motif embedded in histone and non-histone proteins. Interestingly, the CDY chromodomains exhibit different binding preferences for the lysine-methylated ARKS motif in different sequence contexts. Here, we present the structural basis for selective binding of CDY1 to H3K9me3 and preferential binding of CDYL2 to H3tK27me3 over H3K27me3. In addition, we use a CDYL1/2-selective compound, UNC4850, to gain further insight into the molecular mechanisms underlying CDYL2 binding specificity. Our work also provides critical implications that CDYL1b's role in the regulation of neural development is dependent on its recognition of the lysine-methylated ARKS motif.
- Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Tianjin Medical University, Tianjin 300070, China.
Organizational Affiliation: