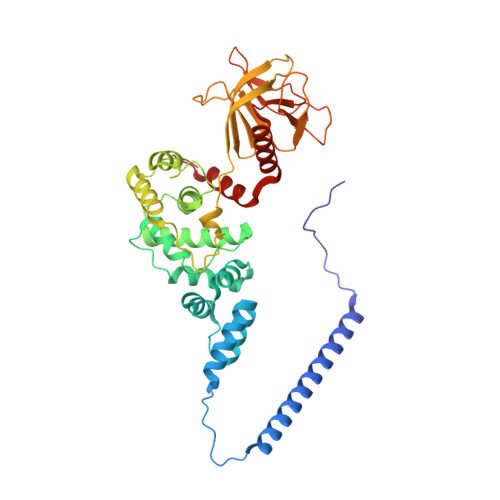
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Das, S., Malaby, A.W., Nawrotek, A., Zhang, W., Zeghouf, M., Maslen, S., Skehel, M., Chakravarthy, S., Irving, T.C., Bilsel, O., Cherfils, J., Lambright, D.G.(2019) Structure 27: 1782
- PubMed: 31601460
- DOI: https://doi.org/10.1016/j.str.2019.09.007
- Primary Citation Related Structures:
6U3E, 6U3G - PubMed Abstract:
Membrane dynamic processes require Arf GTPase activation by guanine nucleotide exchange factors (GEFs) with a Sec7 domain. Cytohesin family Arf GEFs function in signaling and cell migration through Arf GTPase activation on the plasma membrane and endosomes. In this study, the structural organization of two cytohesins (Grp1 and ARNO) was investigated in solution by size exclusion-small angle X-ray scattering and negative stain-electron microscopy and on membranes by dynamic light scattering, hydrogen-deuterium exchange-mass spectrometry and guanosine diphosphate (GDP)/guanosine triphosphate (GTP) exchange assays. The results suggest that cytohesins form elongated dimers with a central coiled coil and membrane-binding pleckstrin-homology (PH) domains at opposite ends. The dimers display significant conformational heterogeneity, with a preference for compact to intermediate conformations. Phosphoinositide-dependent membrane recruitment is mediated by one PH domain at a time and alters the conformational dynamics to prime allosteric activation by Arf-GTP. A structural model for membrane targeting and allosteric activation of full-length cytohesin dimers is discussed.
- Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: