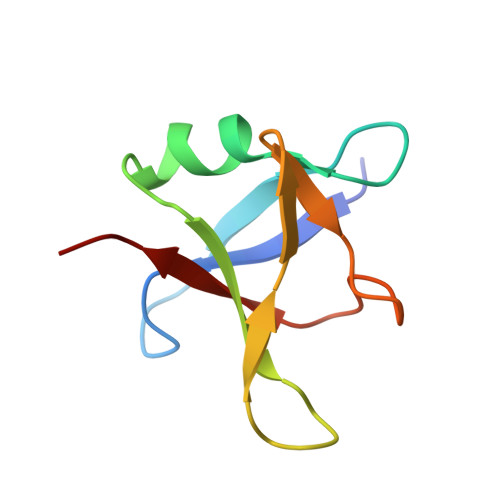
Purification and Structural Characterization of Aggregation-Prone Human TDP-43 Involved in Neurodegenerative Diseases.
Wright, G.S.A., Watanabe, T.F., Amporndanai, K., Plotkin, S.S., Cashman, N.R., Antonyuk, S.V., Hasnain, S.S.(2020) iScience 23: 101159-101159
- PubMed: 32480125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2020.101159
- Primary Citation Related Structures:
6T4B - PubMed Abstract:
Mislocalization, cleavage, and aggregation of the human protein TDP-43 is found in many neurodegenerative diseases. As is the case with many other proteins that are completely or partially structurally disordered, production of full-length recombinant TDP-43 in the quantities necessary for structural characterization has proved difficult. We show that the full-length TDP-43 protein and two truncated N-terminal constructs 1-270 and 1-263 can be heterologously expressed in E. coli. Full-length TDP-43 could be prevented from aggregation during purification using a detergent. Crystals grown from an N-terminal construct (1-270) revealed only the N-terminal domain (residues 1-80) with molecules arranged as parallel spirals with neighboring molecules arranged in head-to-tail fashion. To obtain detergent-free, full-length TDP-43 we mutated all six tryptophan residues to alanine. This provided sufficient soluble protein to collect small-angle X-ray scattering data. Refining relative positions of individual domains and intrinsically disordered regions against this data yielded a model of full-length TDP-43.
- Molecular Biophysics Group, Department of Biochemistry & Systems Biology, Institute of Systems, Molecular and Integrative Biology, Faculty of Health and Life Sciences, Liverpool L69 7ZB, UK.
Organizational Affiliation: