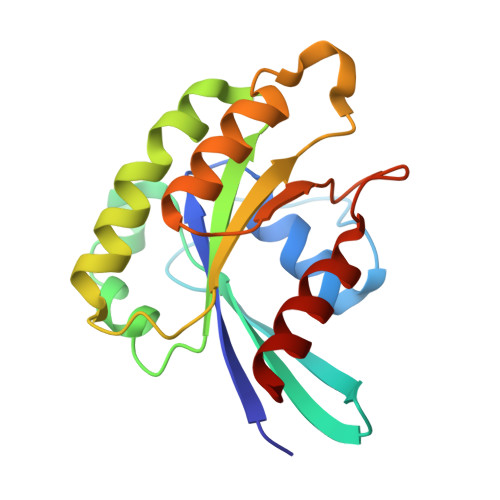
Crystal structure of the Rab33B/Atg16L1 effector complex.
Metje-Sprink, J., Groffmann, J., Neumann, P., Barg-Kues, B., Ficner, R., Kuhnel, K., Schalk, A.M., Binotti, B.(2020) Sci Rep 10: 12956-12956
- PubMed: 32737358 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-69637-0
- Primary Citation Related Structures:
6SUR - PubMed Abstract:
The Atg12-Atg5/Atg16L1 complex is recruited by WIPI2b to the site of autophagosome formation. Atg16L1 is an effector of the Golgi resident GTPase Rab33B. Here we identified a minimal stable complex of murine Rab33B(30-202) Q92L and Atg16L1(153-210). Atg16L1(153-210) comprises the C-terminal part of the Atg16L1 coiled-coil domain. We have determined the crystal structure of the Rab33B Q92L/Atg16L1(153-210) effector complex at 3.47 Å resolution. This structure reveals that two Rab33B molecules bind to the diverging α-helices of the dimeric Atg16L1 coiled-coil domain. We mutated Atg16L1 and Rab33B interface residues and found that they disrupt complex formation in pull-down assays and cellular co-localization studies. The Rab33B binding site of Atg16L1 comprises 20 residues and immediately precedes the WIPI2b binding site. Rab33B mutations that abolish Atg16L binding also abrogate Rab33B association with the Golgi stacks. Atg16L1 mutants that are defective in Rab33B binding still co-localize with WIPI2b in vivo. The close proximity of the Rab33B and WIPI2b binding sites might facilitate the recruitment of Rab33B containing vesicles to provide a source of lipids during autophagosome biogenesis.
- Department of Neurobiology, Max-Planck-Institute for Biophysical Chemistry, 37077, Göttingen, Germany. janina.metje@julius-kuehn.de.
Organizational Affiliation: