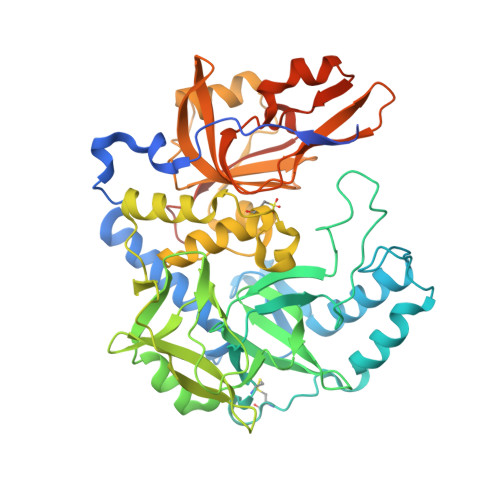
A Secreted NlpC/P60 Endopeptidase from Photobacterium damselae subsp. piscicida Cleaves the Peptidoglycan of Potentially Competing Bacteria.
Lisboa, J., Pereira, C., Rifflet, A., Ayala, J., Terceti, M.S., Barca, A.V., Rodrigues, I., Pereira, P.J.B., Osorio, C.R., Garcia-Del Portillo, F., Gomperts Boneca, I., do Vale, A., Dos Santos, N.M.S.(2021) mSphere 6
- PubMed: 33536321 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mSphere.00736-20
- Primary Citation Related Structures:
6SQX - PubMed Abstract:
Peptidoglycan (PG) is a major component of the bacterial cell wall, forming a mesh-like structure enwrapping the bacteria that is essential for maintaining structural integrity and providing support for anchoring other components of the cell envelope. PG biogenesis is highly dynamic and requires multiple enzymes, including several hydrolases that cleave glycosidic or amide bonds in the PG. This work describes the structural and functional characterization of an NlpC/P60-containing peptidase from Photobacterium damselae subsp. piscicida ( Phdp ), a Gram-negative bacterium that causes high mortality of warm-water marine fish with great impact for the aquaculture industry. PnpA ( P hotobacterium N lpC-like p rotein A ) has a four-domain structure with a hydrophobic and narrow access to the catalytic center and specificity for the γ-d-glutamyl- meso -diaminopimelic acid bond. However, PnpA does not cleave the PG of Phdp or PG of several Gram-negative and Gram-positive bacterial species. Interestingly, it is secreted by the Phdp type II secretion system and degrades the PG of Vibrio anguillarum and Vibrio vulnificus This suggests that PnpA is used by Phdp to gain an advantage over bacteria that compete for the same resources or to obtain nutrients in nutrient-scarce environments. Comparison of the muropeptide composition of PG susceptible and resistant to the catalytic activity of PnpA showed that the global content of muropeptides is similar, suggesting that susceptibility to PnpA is determined by the three-dimensional organization of the muropeptides in the PG. IMPORTANCE Peptidoglycan (PG) is a major component of the bacterial cell wall formed by long chains of two alternating sugars interconnected by short peptides, generating a mesh-like structure that enwraps the bacterial cell. Although PG provides structural integrity and support for anchoring other components of the cell envelope, it is constantly being remodeled through the action of specific enzymes that cleave or join its components. Here, it is shown that Photobacterium damselae subsp. piscicida , a bacterium that causes high mortality in warm-water marine fish, produces PnpA, an enzyme that is secreted into the environment and is able to cleave the PG of potentially competing bacteria, either to gain a competitive advantage and/or to obtain nutrients. The specificity of PnpA for the PG of some bacteria and its inability to cleave others may be explained by differences in the structure of the PG mesh and not by different muropeptide composition.
- Fish Immunology and Vaccinology Group, Instituto de Biologia Molecular e Celular (IBMC), Universidade do Porto, Porto, Portugal johnny.lisboa@ibmc.up.pt nsantos@ibmc.up.pt.
Organizational Affiliation: