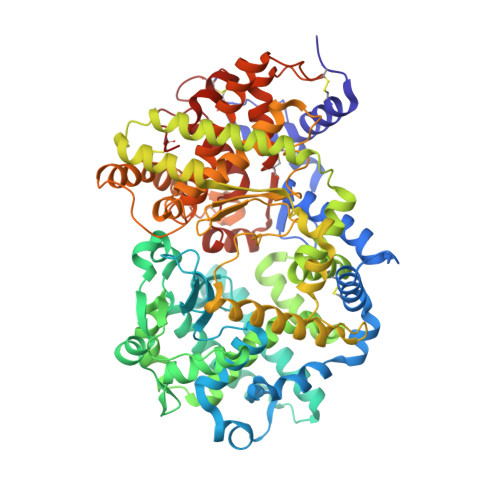
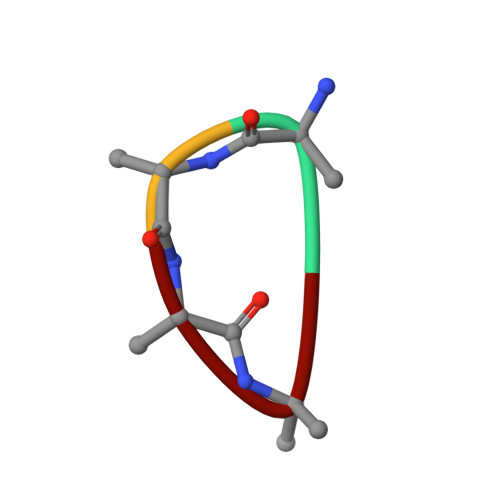
Crystal structure of peptide-bound neprilysin reveals key binding interactions.
Moss, S., Subramanian, V., Acharya, K.R.(2020) FEBS Lett 594: 327-336
- PubMed: 31514225 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13602
- Primary Citation Related Structures:
6SH1, 6SH2 - PubMed Abstract:
Neprilysin (NEP) is a promiscuous zinc metalloprotease with broad substrate specificity and cleaves a remarkable diversity of substrates through endopeptidase action. Two of these - amyloid-β and natriuretic peptides - implicate the enzyme in both Alzheimer's disease and cardiovascular disease, respectively. Here, we report the creation of a catalytically inactive NEP (E584D) to determine the first peptide-bound crystal structure at 2.6 Å resolution. The structure reveals key interactions involved in substrate binding which we have identified to be conserved in other known zinc metalloproteases. In addition, the structure provides evidence for a potential exosite within the central cavity that may play a critical role in substrate positioning. Together, these results contribute to our understanding of the molecular function of NEP.
- Department of Biology and Biochemistry, Claverton Down, University of Bath, UK.
Organizational Affiliation: