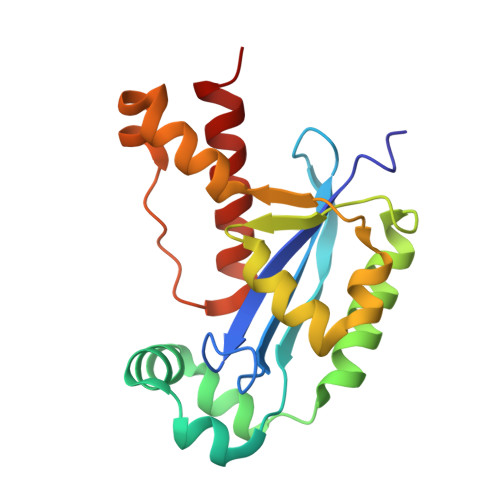
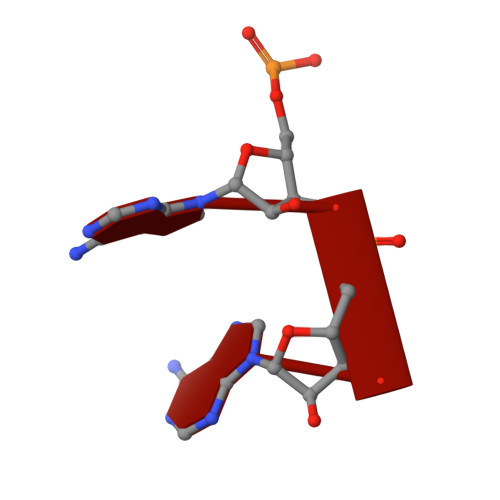
Dinucleotide Degradation by REXO2 Maintains Promoter Specificity in Mammalian Mitochondria.
Nicholls, T.J., Spahr, H., Jiang, S., Siira, S.J., Koolmeister, C., Sharma, S., Kauppila, J.H.K., Jiang, M., Kaever, V., Rackham, O., Chabes, A., Falkenberg, M., Filipovska, A., Larsson, N.G., Gustafsson, C.M.(2019) Mol Cell 76: 784-796.e6
- PubMed: 31588022
- DOI: https://doi.org/10.1016/j.molcel.2019.09.010
- Primary Citation of Related Structures:
6RCI, 6RCL, 6RCN - PubMed Abstract:
Oligoribonucleases are conserved enzymes that degrade short RNA molecules of up to 5 nt in length and are assumed to constitute the final stage of RNA turnover. Here we demonstrate that REXO2 is a specialized dinucleotide-degrading enzyme that shows no preference between RNA and DNA dinucleotide substrates. A heart- and skeletal-muscle-specific knockout mouse displays elevated dinucleotide levels and alterations in gene expression patterns indicative of aberrant dinucleotide-primed transcription initiation. We find that dinucleotides act as potent stimulators of mitochondrial transcription initiation in vitro. Our data demonstrate that increased levels of dinucleotides can be used to initiate transcription, leading to an increase in transcription levels from both mitochondrial promoters and other, nonspecific sequence elements in mitochondrial DNA. Efficient RNA turnover by REXO2 is thus required to maintain promoter specificity and proper regulation of transcription in mammalian mitochondria.
Organizational Affiliation:
Department of Medical Biochemistry and Cell Biology, University of Gothenburg, PO Box 440, Gothenburg 405 30, Sweden.