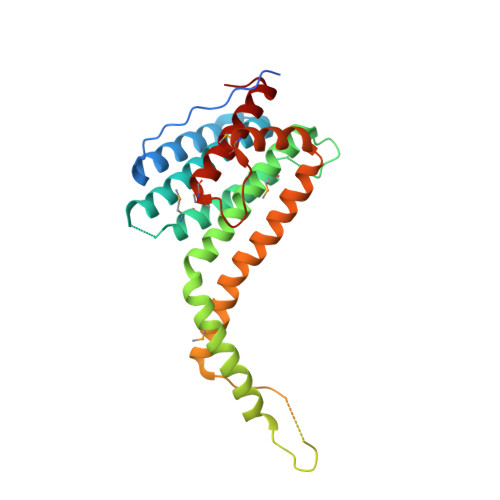
Structure of the super-elongation complex subunit AFF4 C-terminal homology domain reveals requirements for AFF homo- and heterodimerization.
Chen, Y., Cramer, P.(2019) J Biological Chem 294: 10663-10673
- PubMed: 31147444 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.008577
- Primary Citation Related Structures:
6R80 - PubMed Abstract:
AF4/FMR2 family member 4 (AFF4) is the scaffold protein of the multisubunit super-elongation complex, which plays key roles in the release of RNA polymerase II from promoter-proximal pausing and in the transactivation of HIV-1 transcription. AFF4 consists of an intrinsically disordered N-terminal region that interacts with other super-elongation complex subunits and a C-terminal homology domain (CHD) that is conserved among AF4/FMR2 family proteins, including AFF1, AFF2, AFF3, and AFF4. Here, we solved the X-ray crystal structure of the CHD in human AFF4 (AFF4-CHD) to 2.2 Å resolution and characterized its biochemical properties. The structure disclosed that AFF4-CHD folds into a novel domain that consists of eight helices and is distantly related to tetratrico peptide repeat motifs. Our analyses further revealed that AFF4-CHD mediates the formation of an AFF4 homodimer or an AFF1-AFF4 heterodimer. Results from fluorescence anisotropy experiments suggested that AFF4-CHD interacts with both RNA and DNA in vitro Furthermore, we identified a surface loop region in AFF4-CHD as a substrate for the P-TEFb kinase cyclin-dependent kinase 9, which triggers release of polymerase II from promoter-proximal pausing sites. In conclusion, the AFF-CHD structure and biochemical analyses reported here reveal the molecular basis for the homo- and heterodimerization of AFF proteins and implicate the AFF4-CHD in nucleic acid interactions. The high conservation of the CHD among several other proteins suggests that our results are also relevant for understanding other CHD-containing proteins and their dimerization behavior.
- From the Department of Molecular Biology, Max Planck Institute for Biophysical Chemistry, 37077 Göttingen, Germany.
Organizational Affiliation: