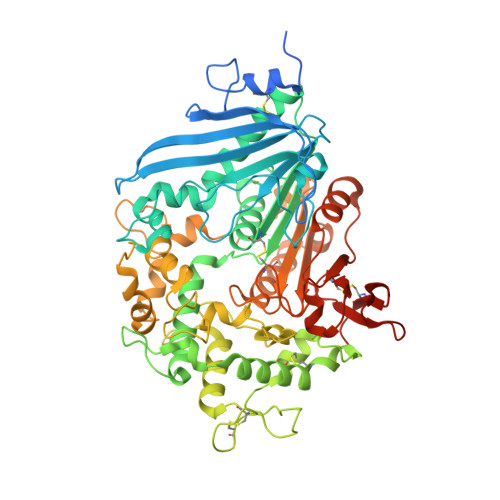
Characterization and engineering of a two-enzyme system for plastics depolymerization.
Knott, B.C., Erickson, E., Allen, M.D., Gado, J.E., Graham, R., Kearns, F.L., Pardo, I., Topuzlu, E., Anderson, J.J., Austin, H.P., Dominick, G., Johnson, C.W., Rorrer, N.A., Szostkiewicz, C.J., Copie, V., Payne, C.M., Woodcock, H.L., Donohoe, B.S., Beckham, G.T., McGeehan, J.E.(2020) Proc Natl Acad Sci U S A 117: 25476-25485
- PubMed: 32989159
- DOI: https://doi.org/10.1073/pnas.2006753117
- Primary Citation Related Structures:
6QZ1, 6QZ2, 6QZ3, 6QZ4 - PubMed Abstract:
Plastics pollution represents a global environmental crisis. In response, microbes are evolving the capacity to utilize synthetic polymers as carbon and energy sources. Recently, Ideonella sakaiensis was reported to secrete a two-enzyme system to deconstruct polyethylene terephthalate (PET) to its constituent monomers. Specifically, the I. sakaiensis PETase depolymerizes PET, liberating soluble products, including mono(2-hydroxyethyl) terephthalate (MHET), which is cleaved to terephthalic acid and ethylene glycol by MHETase. Here, we report a 1.6 Å resolution MHETase structure, illustrating that the MHETase core domain is similar to PETase, capped by a lid domain. Simulations of the catalytic itinerary predict that MHETase follows the canonical two-step serine hydrolase mechanism. Bioinformatics analysis suggests that MHETase evolved from ferulic acid esterases, and two homologous enzymes are shown to exhibit MHET turnover. Analysis of the two homologous enzymes and the MHETase S131G mutant demonstrates the importance of this residue for accommodation of MHET in the active site. We also demonstrate that the MHETase lid is crucial for hydrolysis of MHET and, furthermore, that MHETase does not turnover mono(2-hydroxyethyl)-furanoate or mono(2-hydroxyethyl)-isophthalate. A highly synergistic relationship between PETase and MHETase was observed for the conversion of amorphous PET film to monomers across all nonzero MHETase concentrations tested. Finally, we compare the performance of MHETase:PETase chimeric proteins of varying linker lengths, which all exhibit improved PET and MHET turnover relative to the free enzymes. Together, these results offer insights into the two-enzyme PET depolymerization system and will inform future efforts in the biological deconstruction and upcycling of mixed plastics.
- Renewable Resources and Enabling Sciences Center, National Renewable Energy Laboratory, Golden, CO 80401.
Organizational Affiliation: