Crystal structure of DYRK1A complexed with FC162 inhibitor
Chaikuad, A., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Besson, T., Knapp, S., Structural Genomics Consortium (SGC)To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Dual specificity tyrosine-phosphorylation-regulated kinase 1A | 361 | Homo sapiens | Mutation(s): 0 Gene Names: DYRK1A, DYRK, MNB, MNBH EC: 2.7.12.1 (PDB Primary Data), 2.7.11.23 (UniProt) | 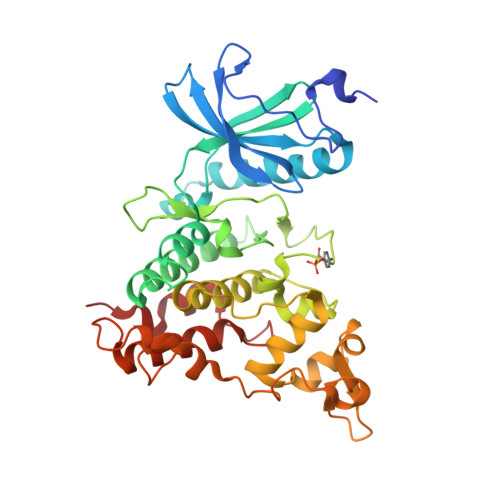 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q13627 GTEx: ENSG00000157540 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q13627 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| JHW Download:Ideal Coordinates CCD File | AA [auth D], J [auth A], R [auth B], W [auth C] | 8-cyclopropyl-2-pyridin-3-yl-[1,3]thiazolo[5,4-f]quinazolin-9-one C17 H12 N4 O S UZKITQOBTXEXSY-UHFFFAOYSA-N |  | ||
| EPE Download:Ideal Coordinates CCD File | V [auth C] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| PG4 Download:Ideal Coordinates CCD File | G [auth A], H [auth A], N [auth B], U [auth C], Z [auth D] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | E [auth A] F [auth A] K [auth B] L [auth B] M [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| DMS Download:Ideal Coordinates CCD File | I [auth A], O [auth B], P [auth B], Q [auth B] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| PTR Query on PTR | A, B, C, D | L-PEPTIDE LINKING | C9 H12 N O6 P |  | TYR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 245.886 | α = 90 |
| b = 65.889 | β = 115.5 |
| c = 148.337 | γ = 90 |
| Software Name | Purpose |
|---|---|
| Aimless | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |