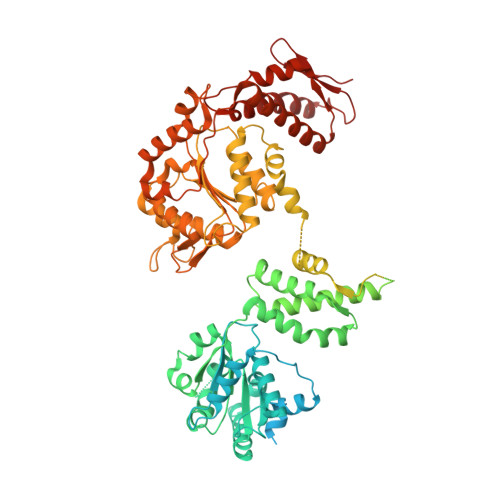
Two-Step Activation Mechanism of the ClpB Disaggregase for Sequential Substrate Threading by the Main ATPase Motor.
Deville, C., Franke, K., Mogk, A., Bukau, B., Saibil, H.R.(2019) Cell Rep 27: 3433-3446.e4
- PubMed: 31216466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2019.05.075
- Primary Citation Related Structures:
6QS4, 6QS6, 6QS7, 6QS8, 6RN2, 6RN3, 6RN4 - PubMed Abstract:
AAA+ proteins form asymmetric hexameric rings that hydrolyze ATP and thread substrate proteins through a central channel via mobile substrate-binding pore loops. Understanding how ATPase and threading activities are regulated and intertwined is key to understanding the AAA+ protein mechanism. We studied the disaggregase ClpB, which contains tandem ATPase domains (AAA1, AAA2) and shifts between low and high ATPase and threading activities. Coiled-coil M-domains repress ClpB activity by encircling the AAA1 ring. Here, we determine the mechanism of ClpB activation by comparing ATPase mechanisms and cryo-EM structures of ClpB wild-type and a constitutively active ClpB M-domain mutant. We show that ClpB activation reduces ATPase cooperativity and induces a sequential mode of ATP hydrolysis in the AAA2 ring, the main ATPase motor. AAA1 and AAA2 rings do not work synchronously but in alternating cycles. This ensures high grip, enabling substrate threading via a processive, rope-climbing mechanism.
- Department of Crystallography, Institute of Structural and Molecular Biology, Birkbeck, University of London, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: