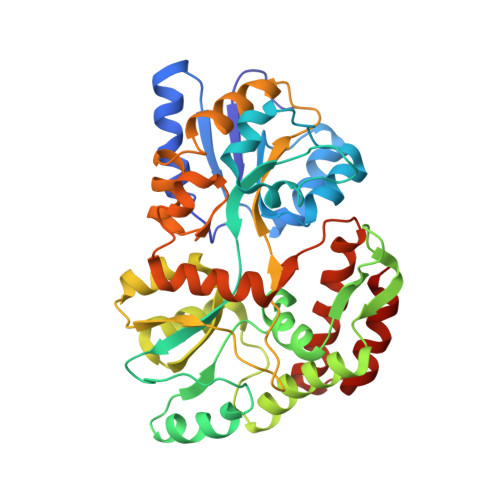
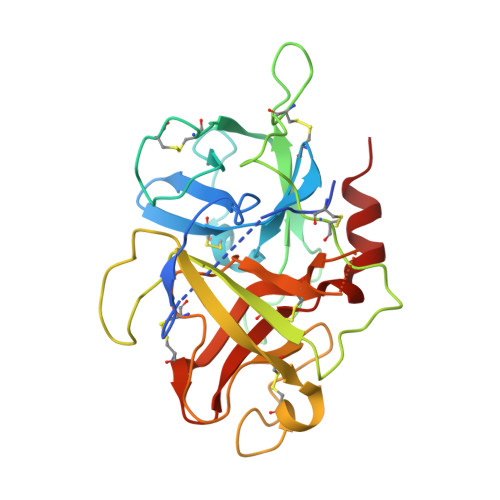
Crystal structures of the recombinant beta-factor XIIa protease with bound Thr-Arg and Pro-Arg substrate mimetics.
Pathak, M., Manna, R., Li, C., Kaira, B.G., Hamad, B.K., Belviso, B.D., Bonturi, C.R., Dreveny, I., Fischer, P.M., Dekker, L.V., Oliva, M.L.V., Emsley, J.(2019) Acta Crystallogr D Struct Biol 75: 578-591
- PubMed: 31205020 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319006910
- Primary Citation Related Structures:
6GT6, 6QF7 - PubMed Abstract:
Coagulation factor XII (FXII) is a key initiator of the contact pathway, which contributes to inflammatory pathways. FXII circulates as a zymogen, which when auto-activated forms factor XIIa (FXIIa). Here, the production of the recombinant FXIIa protease domain (βFXIIa His ) with yields of ∼1-2 mg per litre of insect-cell culture is reported. A second construct utilized an N-terminal maltose-binding protein (MBP) fusion (MBP-βFXIIa His ). Crystal structures were determined of MBP-βFXIIa His in complex with the inhibitor D-Phe-Pro-Arg chloromethyl ketone (PPACK) and of βFXIIa His in isolation. The βFXIIa His structure revealed that the S2 and S1 pockets were occupied by Thr and Arg residues, respectively, from an adjacent molecule in the crystal. The Thr-Arg sequence mimics the P2-P1 FXIIa cleavage-site residues present in the natural substrates prekallikrein and FXII, and Pro-Arg (from PPACK) mimics the factor XI cleavage site. A comparison of the βFXIIa His structure with the available crystal structure of the zymogen-like FXII protease revealed large conformational changes centred around the S1 pocket and an alternate conformation for the 99-loop, Tyr99 and the S2 pocket. Further comparison with activated protease structures of factors IXa and Xa, which also have the Tyr99 residue, reveals that a more open form of the S2 pocket only occurs in the presence of a substrate mimetic. The FXIIa inhibitors EcTI and infestin-4 have Pro-Arg and Phe-Arg P2-P1 sequences, respectively, and the interactions that these inhibitors make with βFXIIa are also described. These structural studies of βFXIIa provide insight into substrate and inhibitor recognition and establish a scaffold for the structure-guided drug design of novel antithrombotic and anti-inflammatory agents.
- Centre for Biomolecular Sciences, School of Pharmacy, University of Nottingham, Nottingham NG7 2RD, England.
Organizational Affiliation: