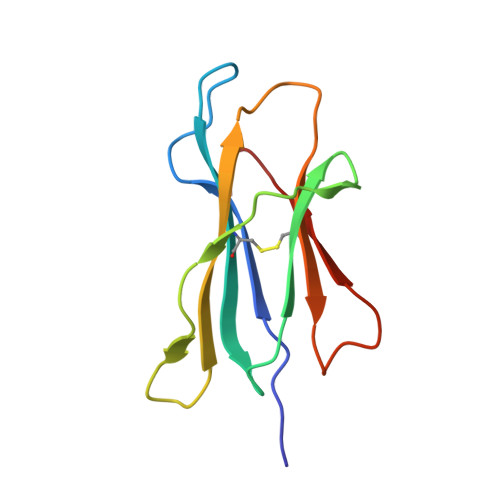
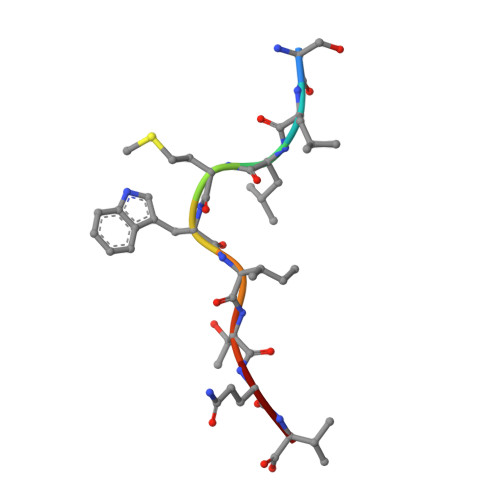
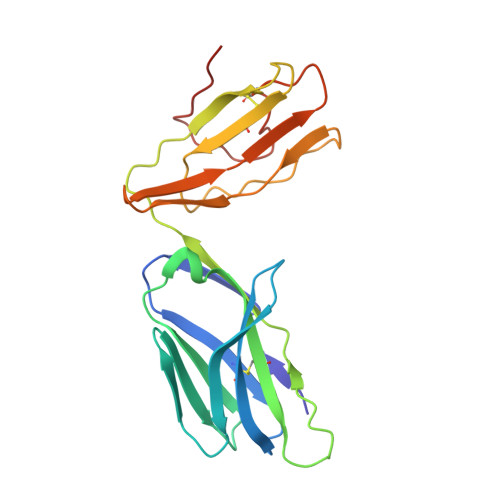
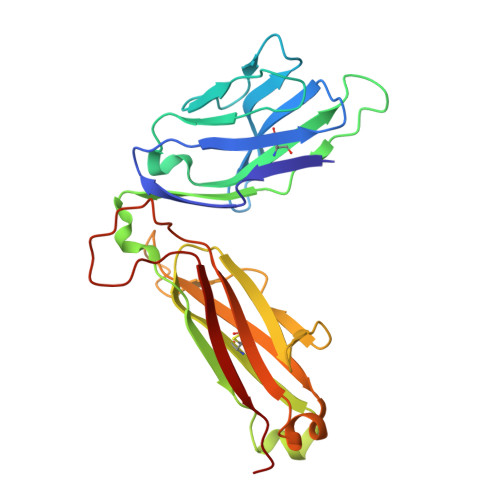
High-throughput peptide-MHC complex generation and kinetic screenings of TCRs with peptide-receptive HLA-A*02:01 molecules.
Moritz, A., Anjanappa, R., Wagner, C., Bunk, S., Hofmann, M., Pszolla, G., Saikia, A., Garcia-Alai, M., Meijers, R., Rammensee, H.G., Springer, S., Maurer, D.(2019) Sci Immunol 4
- PubMed: 31324691 Search on PubMed
- DOI: https://doi.org/10.1126/sciimmunol.aav0860
- Primary Citation Related Structures:
6Q3S - PubMed Abstract:
Major histocompatibility complex (MHC) class I molecules present short peptide ligands on the cell surface for interrogation by cytotoxic CD8 + T cells. MHC class I complexes presenting tumor-associated peptides such as neoantigens represent key targets of cancer immunotherapy approaches currently in development, making them important for efficacy and safety screenings. Without peptide ligand, MHC class I complexes are unstable and decay quickly, making the production of soluble monomers for analytical purposes labor intensive. We have developed a disulfide-stabilized HLA-A*02:01 molecule that is stable without peptide but can form peptide-MHC complexes (pMHCs) with ligands of choice in a one-step loading procedure. We illustrate the similarity between the engineered mutant and the wild-type molecule with respect to affinity of wild-type or affinity-matured T cell receptors (TCRs) and present a crystal structure corroborating the binding kinetics measurements. In addition, we demonstrate a high-throughput binding kinetics measurement platform to analyze the binding characteristics of bispecific TCR (bsTCR) molecules against diverse pMHC libraries produced with the disulfide-stabilized HLA-A*02:01 molecule. We show that bsTCR affinities for pMHCs are indicative of in vitro function and generate a bsTCR binding motif to identify potential off-target interactions in the human proteome. These findings showcase the potential of the platform and the engineered HLA-A*02:01 molecule in the emerging field of pMHC-targeting biologics.
- Department of Immunology, Institute for Cell Biology, University of Tübingen, Tübingen, Germany. andreas.moritz@student.uni-tuebingen.de maurer@immatics.com.
Organizational Affiliation: