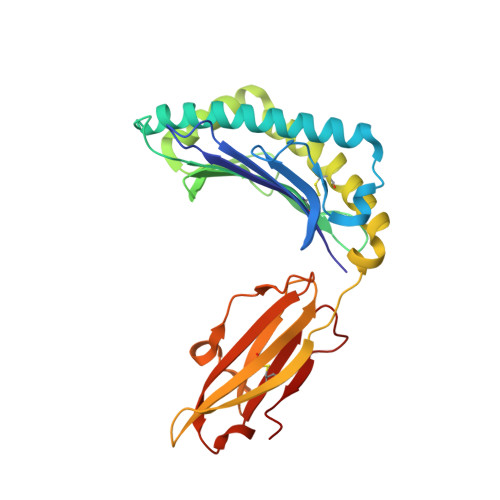
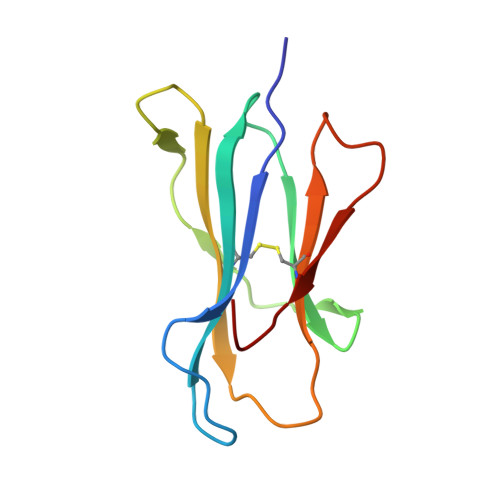
Allelic association with ankylosing spondylitis fails to correlate with human leukocyte antigen B27 homodimer formation.
Lim Kam Sian, T.C.C., Indumathy, S., Halim, H., Greule, A., Cryle, M.J., Bowness, P., Rossjohn, J., Gras, S., Purcell, A.W., Schittenhelm, R.B.(2019) J Biological Chem 294: 20185-20195
- PubMed: 31740583
- DOI: https://doi.org/10.1074/jbc.RA119.010257
- Primary Citation of Related Structures:
6PYJ, 6PYL, 6PYV, 6PYW, 6PZ5 - PubMed Abstract:
Expression of human leukocyte antigen (HLA)-B27 is strongly associated with predisposition toward ankylosing spondylitis (AS) and other spondyloarthropathies. However, the exact involvement of HLA-B27 in disease initiation and progression remains unclear. The homodimer theory, which proposes that HLA-B27 heavy chains aberrantly form homodimers, is a central hypothesis that attempts to explain the role of HLA-B27 in disease pathogenesis. Here, we examined the ability of the eight most prevalent HLA-B27 allotypes (HLA-B*27:02 to HLA-B*27:09) to form homodimers. We observed that HLA-B*27:03, a disease-associated HLA-B27 subtype, showed a significantly reduced ability to form homodimers compared with all other allotypes, including the non-disease-associated/protective allotypes HLA-B*27:06 and HLA-B*27:09. We used X-ray crystallography and site-directed mutagenesis to unravel the molecular and structural mechanisms in HLA-B*27:03 that are responsible for its compromised ability to form homodimers. We show that polymorphism at position 59, which differentiates HLA-B*27:03 from all other allotypes, is responsible for its compromised ability to form homodimers. Indeed, histidine 59 in HLA-B*27:03 leads to a series of local conformational changes that act in concert to reduce the accessibility of the nearby cysteine 67, an essential amino acid residue for the formation of HLA-B27 homodimers. Considered together, the ability of both protective and disease-associated HLA-B27 allotypes to form homodimers and the failure of HLA-B*27:03 to form homodimers challenge the role of HLA-B27 homodimers in AS pathoetiology. Rather, this work implicates other features, such as peptide binding and antigen presentation, as pivotal mechanisms for disease pathogenesis.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: