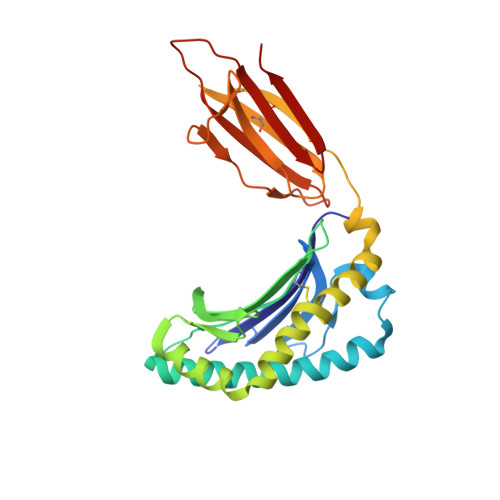
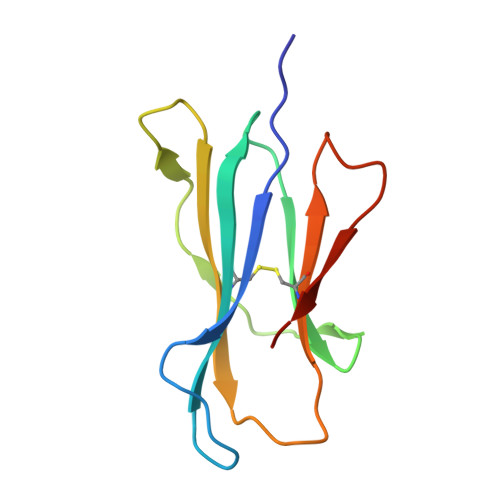
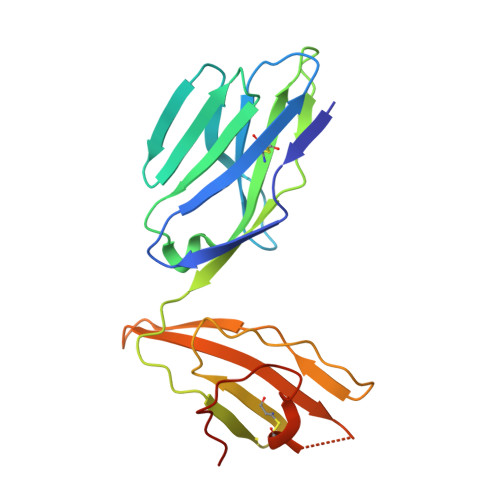
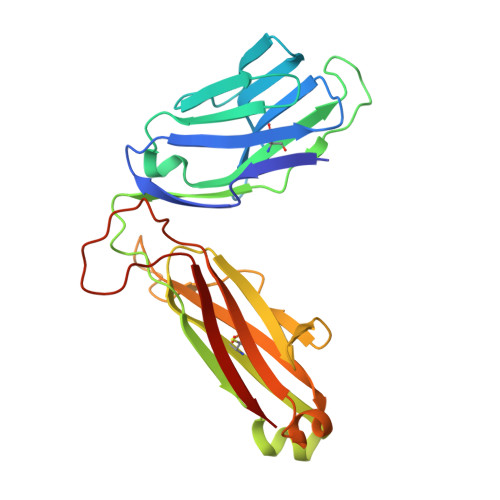
The molecular basis underpinning the potency and specificity of MAIT cell antigens.
Awad, W., Ler, G.J.M., Xu, W., Keller, A.N., Mak, J.Y.W., Lim, X.Y., Liu, L., Eckle, S.B.G., Le Nours, J., McCluskey, J., Corbett, A.J., Fairlie, D.P., Rossjohn, J.(2020) Nat Immunol 21: 400-411
- PubMed: 32123373
- DOI: https://doi.org/10.1038/s41590-020-0616-6
- Primary Citation Related Structures:
6PUC, 6PUD, 6PUE, 6PUF, 6PUG, 6PUH, 6PUI, 6PUJ, 6PUK, 6PUL, 6PUM - PubMed Abstract:
Mucosal-associated invariant T (MAIT) cells are activated by microbial riboflavin-based metabolite antigens when presented by MR1. How modifications to the potent antigen 5-OP-RU affect presentation by MR1 and MAIT cell activation remains unclear. Here we design 20 derivatives, termed altered metabolite ligands (AMLs), to dissect the impact of different antigen components on the human MAIT-MR1 axis. Analysis of 11 crystal structures of MAIT T cell antigen receptor (TCR)-MR1-AML ternary complexes, along with biochemical and functional assays, shows that MR1 cell-surface upregulation is influenced by ribityl and non-ribityl components of the ligand and the hydrophobicity of the MR1-AML interface. The polar ribityl chain of the AML strongly influences MAIT cell activation potency through dynamic compensatory interactions within a MAIT TCR-MR1-AML interaction triad. We define the basis by which the MAIT TCR can differentially recognize AMLs, thereby providing insight into MAIT cell antigen specificity and potency.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: