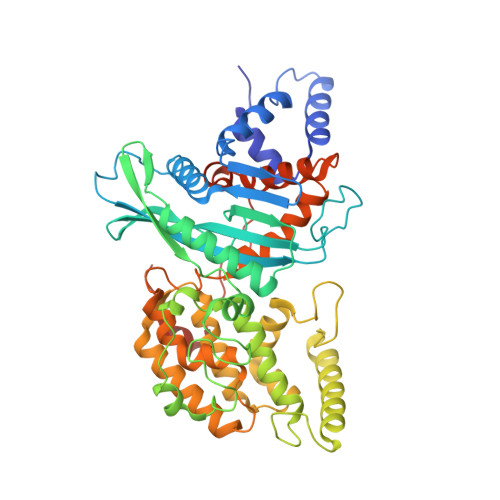
Structures of a RAG-like transposase during cut-and-paste transposition.
Liu, C., Yang, Y., Schatz, D.G.(2019) Nature 575: 540-544
- PubMed: 31723264 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-019-1753-7
- Primary Citation Related Structures:
6PQN, 6PQR, 6PQU, 6PQX, 6PQY, 6PR5 - PubMed Abstract:
Transposons have had a pivotal role in genome evolution 1 and are believed to be the evolutionary progenitors of the RAG1-RAG2 recombinase 2 , an essential component of the adaptive immune system in jawed vertebrates 3 . Here we report one crystal structure and five cryo-electron microscopy structures of Transib 4,5 , a RAG1-like transposase from Helicoverpa zea, that capture the entire transposition process from the apo enzyme to the terminal strand transfer complex with transposon ends covalently joined to target DNA, at resolutions of 3.0-4.6 Å. These structures reveal a butterfly-shaped complex that undergoes two cycles of marked conformational changes in which the 'wings' of the transposase unfurl to bind substrate DNA, close to execute cleavage, open to release the flanking DNA and close again to capture and attack target DNA. Transib possesses unique structural elements that compensate for the absence of a RAG2 partner, including a loop that interacts with the transposition target site and an accordion-like C-terminal tail that elongates and contracts to help to control the opening and closing of the enzyme and assembly of the active site. Our findings reveal the detailed reaction pathway of a eukaryotic cut-and-paste transposase and illuminate some of the earliest steps in the evolution of the RAG recombinase.
- Department of Immunobiology, Yale School of Medicine, New Haven, CT, USA.
Organizational Affiliation: