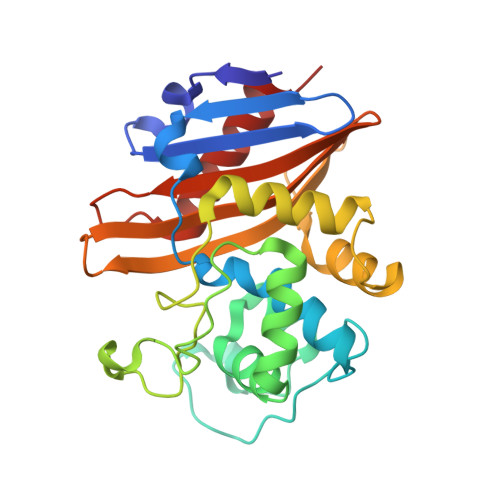
The crystal structures of CDD-1, the intrinsic class D beta-lactamase from the pathogenic Gram-positive bacterium Clostridioides difficile, and its complex with cefotaxime.
Stewart, N.K., Smith, C.A., Toth, M., Stasyuk, A., Vakulenko, S.B.(2019) J Struct Biol 208: 107391-107391
- PubMed: 31550535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2019.09.008
- Primary Citation Related Structures:
6EDM, 6PQC - PubMed Abstract:
Class D β-lactamases, enzymes that degrade β-lactam antibiotics and are widely spread in Gram-negative bacteria, were for a long time not known in Gram-positive organisms. Recently, these enzymes were identified in various non-pathogenic Bacillus species and subsequently in Clostridioides difficile, a major clinical pathogen associated with high morbidity and mortality rates. Comparison of the BPU-1 enzyme from Bacillus pumilus with the CDD-1 and CDD-2 enzymes from C. difficile demonstrated that the latter enzymes have broadened their substrate profile to efficiently hydrolyze the expanded-spectrum methoxyimino cephalosporins, cefotaxime and ceftriaxone. These two antibiotics are major contributors to the development of C. difficile infection, as they suppress sensitive bacterial microflora in the gut but fail to kill the pathogen which is highly resistant to these drugs. To gain insight into the structural features that contribute to the expansion of the substrate profile of CDD enzymes compared to BPU-1, we solved the crystal structures of CDD-1 and its complex with cefotaxime. Comparison of CDD-1 structures with those of class D enzymes from Gram-negative bacteria showed that in the cefotaxime-CDD-1 complex, the antibiotic is bound in a substantially different mode due to structural differences in the enzymes' active sites. We also found that CDD-1 has a uniquely long Ω-loop when compared to all other class D β-lactamases. This Ω-loop extension allows it to engage in hydrogen bonding with the acylated cefotaxime, thus providing additional stabilizing interactions with the substrate which could be responsible for the high catalytic activity of the enzyme for expanded-spectrum cephalosporins.
- Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, IN, USA.
Organizational Affiliation: