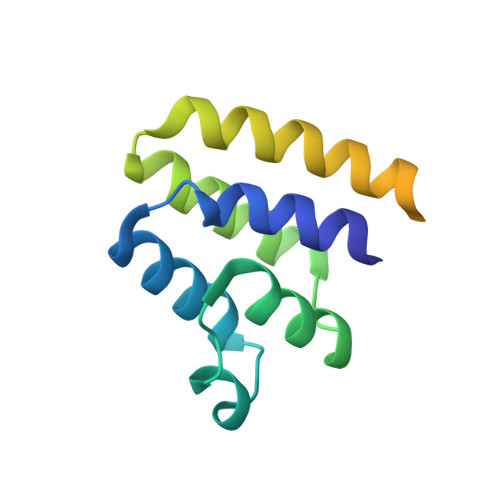
PEA-15 C-Terminal Tail Allosterically Modulates Death-Effector Domain Conformation and Facilitates Protein-Protein Interactions.
Crespo-Flores, S.L., Cabezas, A., Hassan, S., Wei, Y.(2019) Int J Mol Sci 20
- PubMed: 31284641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms20133335
- Primary Citation Related Structures:
6P6B, 6P6C - PubMed Abstract:
Phosphoprotein enriched in astrocytes, 15 kDa (PEA-15) exerts its regulatory roles on several critical cellular pathways through protein-protein interactions depending on its phosphorylation states. It can either inhibit the extracellular signal-regulated kinase (ERK) activities when it is dephosphorylated or block the assembly of death-inducing signaling complex (DISC) and the subsequent activation of apoptotic initiator, caspase-8, when it is phosphorylated. Due to the important roles of PEA-15 in regulating these pathways that lead to opposite cellular outcomes (cell proliferation vs. cell death), we proposed a phosphostasis (phosphorylation homeostasis) model, in which the phosphorylation states of the protein are vigorously controlled and regulated to maintain a delicate balance. The phosphostasis gives rise to the protective cellular functions of PEA-15 to preserve optimum cellular conditions. In this article, using advanced multidimensional nuclear magnetic resonance (NMR) techniques combined with a novel chemical shift (CS)-Rosetta algorithm for de novo protein structural determination, we report a novel conformation of PEA-15 death-effector domain (DED) upon interacting with ERK2. This new conformation is modulated by the irregularly structured C-terminal tail when it first recognizes and binds to ERK2 at the d-peptide recruitment site (DRS) in an allosteric manner, and is facilitated by the rearrangement of the surface electrostatic and hydrogen-bonding interactions on the DED. In this ERK2-bound conformation, three of the six helices (α2, α3, and α4) comprising the DED reorient substantially in comparison to the free-form structure, exposing key residues on the other three helices that directly interact with ERK2 at the DEF-docking site (docking site for ERK, FxF) and the activation loop. Additionally, we provide evidence that the phosphorylation of the C-terminal tail leads to a distinct conformation of DED, allowing efficient interactions with Fas-associated death domain (FADD) protein at the DISC. Our results substantiate the allosteric regulatory roles of the C-terminal tail in modulating DED conformation and facilitating protein-protein interactions of PEA-15.
- Department of Chemistry, New Jersey City University, Jersey City, NJ 07305-1596, USA.
Organizational Affiliation: