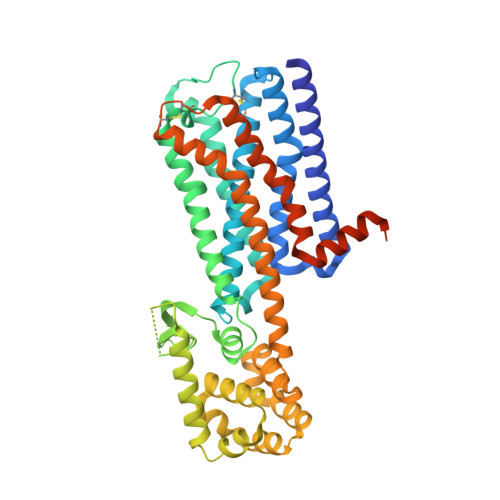
Crystal structure of the M5muscarinic acetylcholine receptor.
Vuckovic, Z., Gentry, P.R., Berizzi, A.E., Hirata, K., Varghese, S., Thompson, G., van der Westhuizen, E.T., Burger, W.A.C., Rahmani, R., Valant, C., Langmead, C.J., Lindsley, C.W., Baell, J.B., Tobin, A.B., Sexton, P.M., Christopoulos, A., Thal, D.M.(2019) Proc Natl Acad Sci U S A 116: 26001-26007
- PubMed: 31772027 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1914446116
- Primary Citation Related Structures:
6OL9 - PubMed Abstract:
The human M 5 muscarinic acetylcholine receptor (mAChR) has recently emerged as an exciting therapeutic target for treating a range of disorders, including drug addiction. However, a lack of structural information for this receptor subtype has limited further drug development and validation. Here we report a high-resolution crystal structure of the human M 5 mAChR bound to the clinically used inverse agonist, tiotropium. This structure allowed for a comparison across all 5 mAChR family members that revealed important differences in both orthosteric and allosteric sites that could inform the rational design of selective ligands. These structural studies, together with chimeric swaps between the extracellular regions of the M 2 and M 5 mAChRs, provided structural insight into kinetic selectivity, where ligands show differential residency times between related family members. Collectively, our study provides important insights into the nature of orthosteric and allosteric ligand interaction across the mAChR family that could be exploited for the design of selective drugs.
- Drug Discovery Biology, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, VIC 3052, Australia.
Organizational Affiliation: