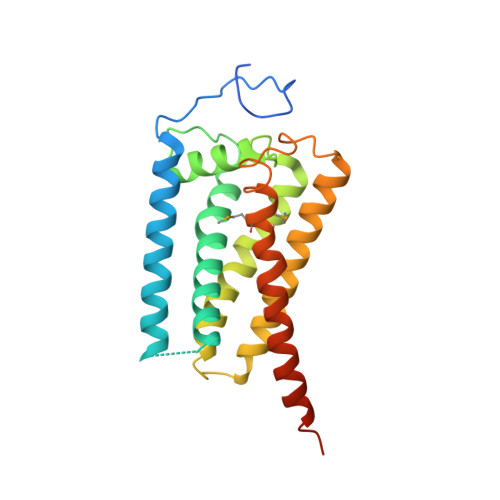
Structural basis for human sterol isomerase in cholesterol biosynthesis and multidrug recognition.
Long, T., Hassan, A., Thompson, B.M., McDonald, J.G., Wang, J., Li, X.(2019) Nat Commun 10: 2452-2452
- PubMed: 31165728 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10279-w
- Primary Citation Related Structures:
6OHT, 6OHU - PubMed Abstract:
3-β-hydroxysteroid-Δ 8 , Δ 7 -isomerase, known as Emopamil-Binding Protein (EBP), is an endoplasmic reticulum membrane protein involved in cholesterol biosynthesis, autophagy, oligodendrocyte formation. The mutation on EBP can cause Conradi-Hunermann syndrome, an inborn error. Interestingly, EBP binds an abundance of structurally diverse pharmacologically active compounds, causing drug resistance. Here, we report two crystal structures of human EBP, one in complex with the anti-breast cancer drug tamoxifen and the other in complex with the cholesterol biosynthesis inhibitor U18666A. EBP adopts an unreported fold involving five transmembrane-helices (TMs) that creates a membrane cavity presenting a pharmacological binding site that accommodates multiple different ligands. The compounds exploit their positively-charged amine group to mimic the carbocationic sterol intermediate. Mutagenesis studies on specific residues abolish the isomerase activity and decrease the multidrug binding capacity. This work reveals the catalytic mechanism of EBP-mediated isomerization in cholesterol biosynthesis and how this protein may act as a multi-drug binder.
- Department of Molecular Genetics, University of Texas Southwestern Medical Center, Dallas, TX, 75390, USA.
Organizational Affiliation: