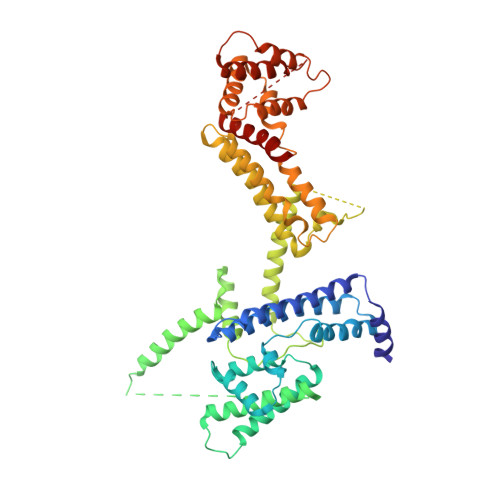
Structure of the Helicobacter pylori Cag type IV secretion system.
Chung, J.M., Sheedlo, M.J., Campbell, A.M., Sawhney, N., Frick-Cheng, A.E., Lacy, D.B., Cover, T.L., Ohi, M.D.(2019) Elife 8
- PubMed: 31210639 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.47644
- Primary Citation Related Structures:
6ODI, 6ODJ, 6OEE, 6OEF, 6OEG, 6OEH - PubMed Abstract:
Bacterial type IV secretion systems (T4SSs) are molecular machines that can mediate interbacterial DNA transfer through conjugation and delivery of effector molecules into host cells. The Helicobacter pylori Cag T4SS translocates CagA, a bacterial oncoprotein, into gastric cells, contributing to gastric cancer pathogenesis. We report the structure of a membrane-spanning Cag T4SS assembly, which we describe as three sub-assemblies: a 14-fold symmetric outer membrane core complex (OMCC), 17-fold symmetric periplasmic ring complex (PRC), and central stalk. Features that differ markedly from those of prototypical T4SSs include an expanded OMCC and unexpected symmetry mismatch between the OMCC and PRC. This structure is one of the largest bacterial secretion system assemblies ever reported and illustrates the remarkable structural diversity that exists among bacterial T4SSs.
- Life Sciences Institute, University of Michigan, Ann Arbor, United States.
Organizational Affiliation: