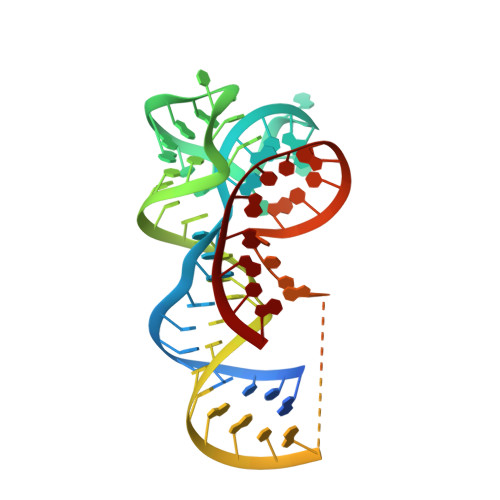
Co-crystal structure of the Fusobacterium ulcerans ZTP riboswitch using an X-ray free-electron laser.
Jones, C., Tran, B., Conrad, C., Stagno, J., Trachman 3rd, R., Fischer, P., Meents, A., Ferre-D'Amare, A.(2019) Acta Crystallogr F Struct Biol Commun 75: 496-500
- PubMed: 31282869 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19008549
- Primary Citation Related Structures:
6OD9 - PubMed Abstract:
Riboswitches are conformationally dynamic RNAs that regulate gene expression by binding specific small molecules. ZTP riboswitches bind the purine-biosynthetic intermediate 5-aminoimidazole-4-carboxamide riboside 5'-monophosphate (ZMP) and its triphosphorylated form (ZTP). Ligand binding to this riboswitch ultimately upregulates genes involved in folate and purine metabolism. Using an X-ray free-electron laser (XFEL), the room-temperature structure of the Fusobacterium ulcerans ZTP riboswitch bound to ZMP has now been determined at 4.1 Å resolution. This model, which was refined against a data set from ∼750 diffraction images (each from a single crystal), was found to be consistent with that previously obtained from data collected at 100 K using conventional synchrotron X-radiation. These experiments demonstrate the feasibility of time-resolved XFEL experiments to understand how the ZTP riboswitch accommodates cognate ligand binding.
- Biochemistry and Biophysics Center, National Heart, Lung and Blood Institute, 50 South Drive, MSC 8012, Bethesda, MD 20892, USA.
Organizational Affiliation: