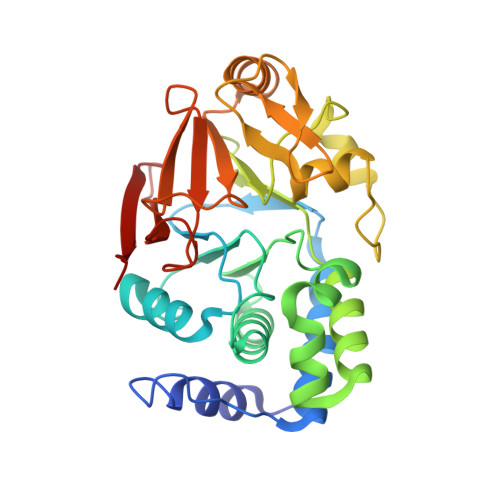
SDS22 selectively recognizes and traps metal-deficient inactive PP1.
Choy, M.S., Moon, T.M., Ravindran, R., Bray, J.A., Robinson, L.C., Archuleta, T.L., Shi, W., Peti, W., Tatchell, K., Page, R.(2019) Proc Natl Acad Sci U S A 116: 20472-20481
- PubMed: 31548429 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1908718116
- Primary Citation Related Structures:
6OBN, 6OBP, 6OBQ, 6OBR, 6OBS, 6OBU - PubMed Abstract:
The metalloenzyme protein phosphatase 1 (PP1), which is responsible for ≥50% of all dephosphorylation reactions, is regulated by scores of regulatory proteins, including the highly conserved SDS22 protein. SDS22 has numerous diverse functions, surprisingly acting as both a PP1 inhibitor and as an activator. Here, we integrate cellular, biophysical, and crystallographic studies to address this conundrum. We discovered that SDS22 selectively binds a unique conformation of PP1 that contains a single metal (M2) at its active site, i.e., SDS22 traps metal-deficient inactive PP1. Furthermore, we showed that SDS22 dissociation is accompanied by a second metal (M1) being loaded into PP1, as free metal cannot dissociate the complex and M1-deficient mutants remain constitutively trapped by SDS22. Together, our findings reveal that M1 metal loading and loss are essential for PP1 regulation in cells, which has broad implications for PP1 maturation, activity, and holoenzyme subunit exchange.
- Department of Chemistry and Biochemistry, University of Arizona, Tucson, AZ 85721.
Organizational Affiliation: