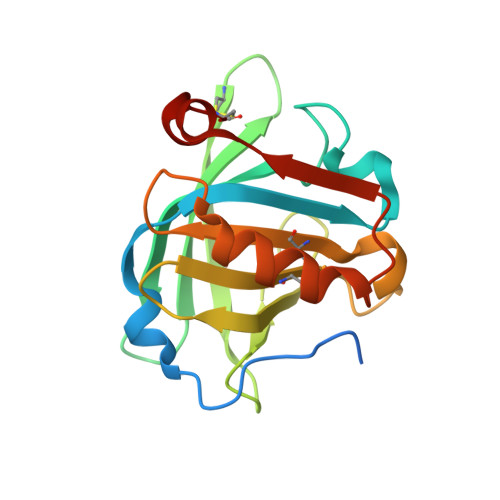
The structure of bovine beta-lactoglobulin in crystals grown at pH 3.8 exhibiting novel threefold twinning.
Yeates, T.O., McPherson, A.(2019) Acta Crystallogr F Struct Biol Commun 75: 640-645
- PubMed: 31584012 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1901224X
- Primary Citation Related Structures:
6NKQ - PubMed Abstract:
Bovine β-lactoglobulin was crystallized from 3 M NaCl buffered at pH 3.8 with sodium citrate as thick hexagonal prisms of greater than 1 mm in edge length. Analyses of the X-ray diffraction intensities using three different current algorithms were unanimous in specifying the space group to be P6 3 22, with unit-cell dimensions a = b = 75.47, c = 140.79 Å. No progress could be made, however, towards an acceptable solution by molecular replacement using this symmetry. In the end, it was found that the true space group was C222 1 , a subgroup of P6 3 22, with a = 65.89, b = 114.12, c = 140.51 Å, with the apparent 622 symmetry arising from an unusual threefold or tritohedral twinning. An assembly based on a model of the protein in another crystal form (PDB entry 1beb) containing three molecules in the asymmetric unit was refined to 2.3 Å resolution with a final R factor of 0.23 and R free of 0.26. NCS restraints were maintained throughout. For the most part, the molecules found in this crystal form are virtually the same as in PDB entry 1beb, although there are numerous local variations, particularly in loop elements, rotamer conformation differences and some alterations, including additions, at the termini.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: