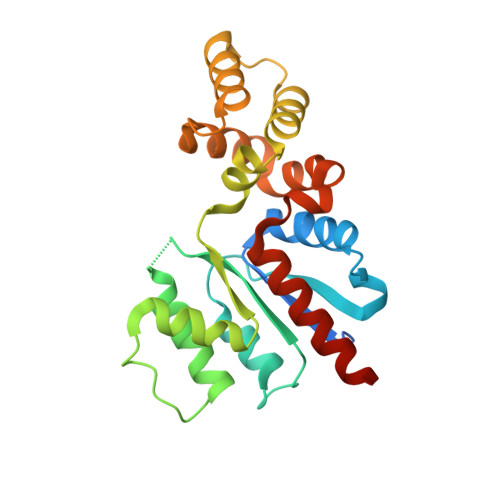
Structural characterization of Class 2 OLD family nucleases supports a two-metal catalysis mechanism for cleavage.
Schiltz, C.J., Lee, A., Partlow, E.A., Hosford, C.J., Chappie, J.S.(2019) Nucleic Acids Res 47: 9448-9463
- PubMed: 31400118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz703
- Primary Citation Related Structures:
6NJV, 6NJW, 6NJX, 6NK8 - PubMed Abstract:
Overcoming lysogenization defect (OLD) proteins constitute a family of uncharacterized nucleases present in bacteria, archaea, and some viruses. These enzymes contain an N-terminal ATPase domain and a C-terminal Toprim domain common amongst replication, recombination, and repair proteins. The in vivo activities of OLD proteins remain poorly understood and no definitive structural information exists. Here we identify and define two classes of OLD proteins based on differences in gene neighborhood and amino acid sequence conservation and present the crystal structures of the catalytic C-terminal regions from the Burkholderia pseudomallei and Xanthamonas campestris p.v. campestris Class 2 OLD proteins at 2.24 Å and 1.86 Å resolution respectively. The structures reveal a two-domain architecture containing a Toprim domain with altered architecture and a unique helical domain. Conserved side chains contributed by both domains coordinate two bound magnesium ions in the active site of B. pseudomallei OLD in a geometry that supports a two-metal catalysis mechanism for cleavage. The spatial organization of these domains additionally suggests a novel mode of DNA binding that is distinct from other Toprim containing proteins. Together, these findings define the fundamental structural properties of the OLD family catalytic core and the underlying mechanism controlling nuclease activity.
- Department of Molecular Medicine, Cornell University, Ithaca, NY 14853, USA.
Organizational Affiliation: