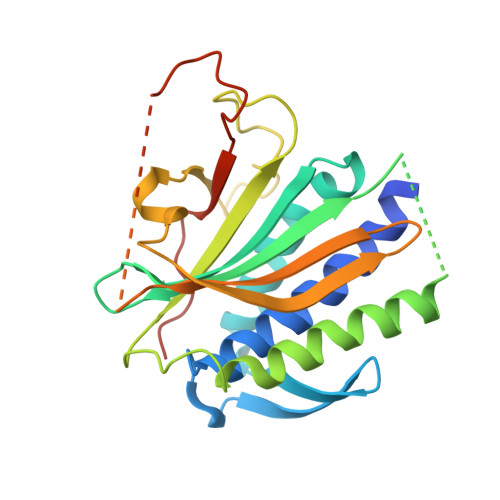
REV7 has a dynamic adaptor region to accommodate small GTPase RAN/ShigellaIpaB ligands, and its activity is regulated by the RanGTP/GDP switch.
Wang, X., Pernicone, N., Pertz, L., Hua, D., Zhang, T., Listovsky, T., Xie, W.(2019) J Biological Chem 294: 15733-15742
- PubMed: 31484720 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.010123
- Primary Citation Related Structures:
6KEA, 6NIF - PubMed Abstract:
REV7, also termed mitotic arrest-deficient 2-like 2 (MAD2L2 or MAD2B), acts as an interaction module in a broad array of cellular pathways, including translesion DNA synthesis, cell cycle control, and nonhomologous end joining. Numerous REV7 binding partners have been identified, including the human small GTPase Ras-associated nuclear protein (RAN), which acts as a potential upstream regulator of REV7. Notably, the Shigella invasin IpaB hijacks REV7 to disrupt cell cycle control to prevent intestinal epithelial cell renewal and facilitate bacterial colonization. However, the structural details of the REV7-RAN and REV7-IpaB interactions are mostly unknown. Here, using fusion protein and rigid maltose-binding protein tagging strategies, we determined the crystal structures of these two complexes at 2.00-2.35 Å resolutions. The structures revealed that both RAN and IpaB fragments bind the "safety belt" region of REV7, inducing rearrangement of the C-terminal β-sheet region of REV7, conserved among REV7-related complexes. Of note, the REV7-binding motifs of RAN and IpaB each displayed some unique interactions with REV7 despite sharing consensus residues. Structural alignments revealed that REV7 has an adaptor region within the safety belt region that can rearrange secondary structures to fit a variety of different ligands. Our structural and biochemical results further indicated that REV7 preferentially binds GTP-bound RAN, implying that a GTP/GDP-bound transition of RAN may serve as the molecular switch that controls REV7's activity. These results provide insights into the regulatory mechanism of REV7 in cell cycle control, which may help with the development of small-molecule inhibitors that target REV7 activity.
- School of Life Sciences, Tianjin University, Tianjin 300072, China.
Organizational Affiliation: