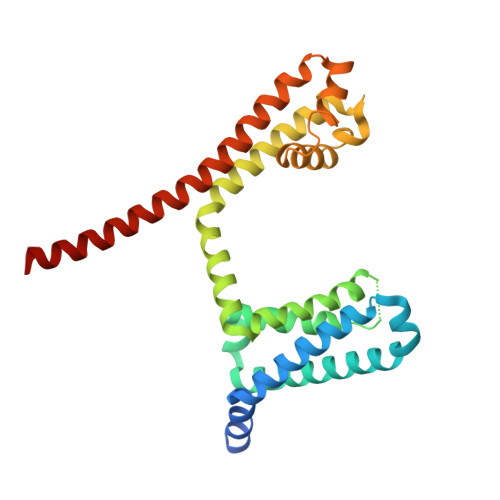
Molecular dissection of multiphase inactivation of the bacterial sodium channel NaVAb.
Gamal El-Din, T.M., Lenaeus, M.J., Ramanadane, K., Zheng, N., Catterall, W.A.(2019) J Gen Physiol 151: 174-185
- PubMed: 30510035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1085/jgp.201711884
- Primary Citation Related Structures:
6MWA, 6MWB, 6MWD, 6MWG - PubMed Abstract:
Homotetrameric bacterial voltage-gated sodium channels share major biophysical features with their more complex eukaryotic counterparts, including a slow-inactivation mechanism that reduces ion-conductance activity during prolonged depolarization through conformational changes in the pore. The bacterial sodium channel Na V Ab activates at very negative membrane potentials and inactivates through a multiphase slow-inactivation mechanism. Early voltage-dependent inactivation during one depolarization is followed by late use-dependent inactivation during repetitive depolarization. Mutations that change the molecular volume of Thr206 in the pore-lining S6 segment can enhance or strongly block early voltage-dependent inactivation, suggesting that this residue serves as a molecular hub controlling the coupling of activation to inactivation. In contrast, truncation of the C-terminal tail enhances the early phase of inactivation yet completely blocks late use-dependent inactivation. Determination of the structure of a C-terminal tail truncation mutant and molecular modeling of conformational changes at Thr206 and the S6 activation gate led to a two-step model of these gating processes. First, bending of the S6 segment, local protein interactions dependent on the size of Thr206, and exchange of hydrogen-bonding partners at the level of Thr206 trigger pore opening followed by the early phase of voltage-dependent inactivation. Thereafter, conformational changes in the C-terminal tail lead to late use-dependent inactivation. These results have important implications for the sequence of conformational changes that lead to multiphase inactivation of Na V Ab and other sodium channels.
- Department of Pharmacology, University of Washington, Seattle, WA.
Organizational Affiliation: