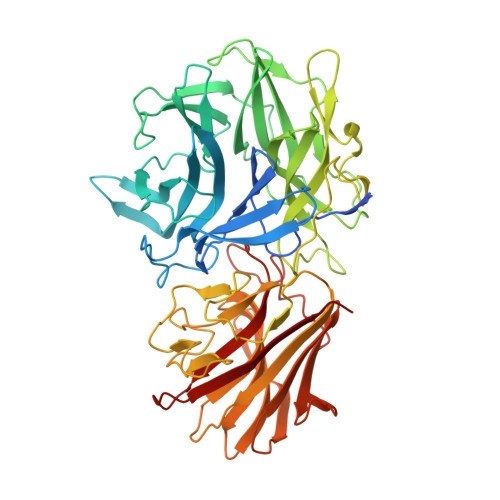
Structure-guided design combined with evolutionary diversity led to the discovery of the xylose-releasing exo-xylanase activity in the glycoside hydrolase family 43.
Zanphorlin, L.M., de Morais, M.A.B., Diogo, J.A., Domingues, M.N., de Souza, F.H.M., Ruller, R., Murakami, M.T.(2019) Biotechnol Bioeng 116: 734-744
- PubMed: 30556897 Search on PubMed
- DOI: https://doi.org/10.1002/bit.26899
- Primary Citation Related Structures:
6MS2, 6MS3 - PubMed Abstract:
Rational design is an important tool for sculpting functional and stability properties of proteins and its potential can be much magnified when combined with in vitro and natural evolutionary diversity. Herein, we report the structure-guided design of a xylose-releasing exo-β-1,4-xylanase from an inactive member of glycoside hydrolase family 43 (GH43). Structural analysis revealed a nonconserved substitution (Lys 247 ) that results in the disruption of the hydrogen bond network that supports catalysis. The mutation of this residue to a conserved serine restored the catalytic activity and crystal structure elucidation of the mutant confirmed the recovery of the proper orientation of the catalytically relevant histidine. Interestingly, the tailored enzyme can cleave both xylooligosaccharides and xylan, releasing xylose as the main product, being the first xylose-releasing exo-β-1,4-xylanase reported in the GH43 family. This enzyme presents a unique active-site topology when compared with closely related β-xylosidases, which is the absence of a hydrophobic barrier at the positive-subsite region, allowing the accommodation of long substrates. Therefore, the combination of rational design for catalytic activation along with naturally occurring differences in the substrate binding interface led to the discovery of a novel activity within the GH43 family. In addition, these results demonstrate the importance of solvation of the β-propeller hollow for GH43 catalytic function and expand our mechanistic understanding about the diverse modes of action of GH43 members, a key and polyspecific carbohydrate-active enzyme family abundant in most plant cell-wall-degrading microorganisms.
- Brazilian Bioethanol Science and Technology Laboratory, National Center for Research in Energy and Materials, Campinas, São Paulo, Brazil.
Organizational Affiliation: