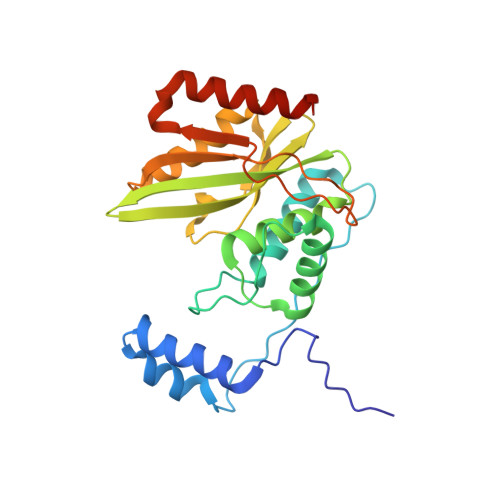
The crystal structure of the tetrameric human vasohibin-1-SVBP complex reveals a variable arm region within the structural core.
Ikeda, A., Urata, S., Ando, T., Suzuki, Y., Sato, Y., Nishino, T.(2020) Acta Crystallogr D Struct Biol 76: 993-1000
- PubMed: 33021501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798320011298
- Primary Citation Related Structures:
6LPG - PubMed Abstract:
Vasohibins regulate angiogenesis, tumor growth, metastasis and neuronal differentiation. They form a complex with small vasohibin-binding protein (SVBP) and show tubulin tyrosine carboxypeptidase activity. Recent crystal structure determinations of vasohibin-SVBP complexes have provided a molecular basis for complex formation, substrate binding and catalytic activity. However, the regulatory mechanism and dynamics of the complex remain elusive. Here, the crystal structure of the VASH1-SVBP complex and a molecular-dynamics simulation study are reported. The overall structure of the complex was similar to previously reported structures. Importantly, however, the structure revealed a domain-swapped heterotetramer that was formed between twofold symmetry-related molecules. This heterotetramerization was stabilized by the mutual exchange of ten conserved N-terminal residues from the VASH1 structural core, which was intramolecular in other structures. Interestingly, a comparison of this region with previously reported structures revealed that the patterns of hydrogen bonding and hydrophobic interactions vary. In the molecular-dynamics simulations, differences were found between the heterotetramer and heterodimer, where the fluctuation of the N-terminal region in the heterotetramer was suppressed. Thus, heterotetramer formation and flexibility of the N-terminal region may be important for enzyme activity and regulation.
- Department of Biological Science and Technology, Tokyo University of Science, 6-3-1 Niijyuku, Katsushika-ku, Tokyo 125-8585, Japan.
Organizational Affiliation: