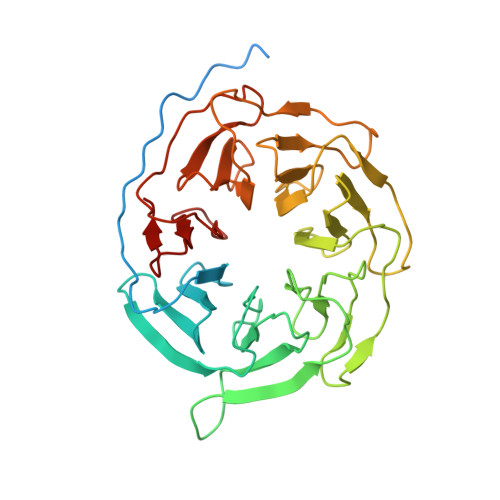
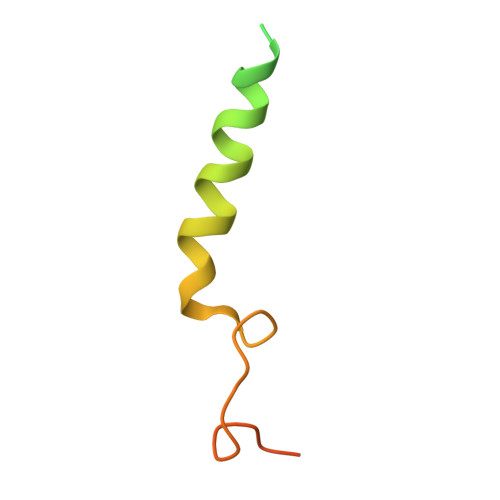
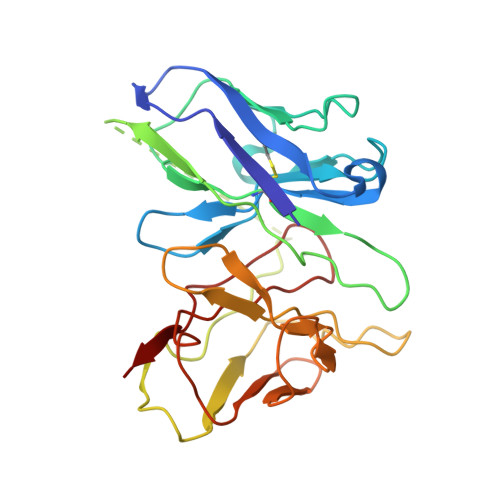
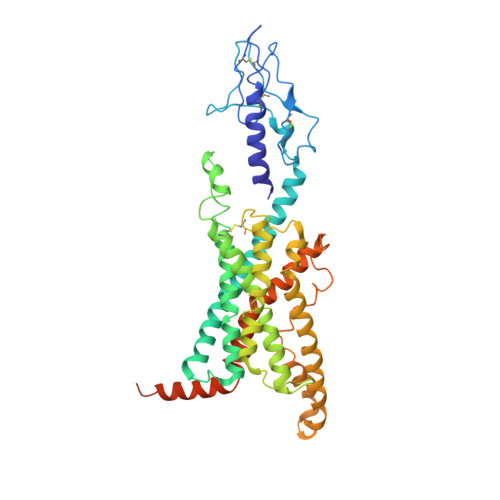
Structural basis of Gsand Girecognition by the human glucagon receptor.
Qiao, A., Han, S., Li, X., Li, Z., Zhao, P., Dai, A., Chang, R., Tai, L., Tan, Q., Chu, X., Ma, L., Thorsen, T.S., Reedtz-Runge, S., Yang, D., Wang, M.W., Sexton, P.M., Wootten, D., Sun, F., Zhao, Q., Wu, B.(2020) Science 367: 1346-1352
- PubMed: 32193322 Search on PubMed
- DOI: https://doi.org/10.1126/science.aaz5346
- Primary Citation Related Structures:
6LMK, 6LML - PubMed Abstract:
Class B G protein-coupled receptors, an important class of therapeutic targets, signal mainly through the G s class of heterotrimeric G proteins, although they do display some promiscuity in G protein binding. Using cryo-electron microscopy, we determined the structures of the human glucagon receptor (GCGR) bound to glucagon and distinct classes of heterotrimeric G proteins, G s or G i1 These two structures adopt a similar open binding cavity to accommodate G s and G i1 The G s binding selectivity of GCGR is explained by a larger interaction interface, but there are specific interactions that affect G i more than G s binding. Conformational differences in the receptor intracellular loops were found to be key selectivity determinants. These distinctions in transducer engagement were supported by mutagenesis and functional studies.
- CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China.
Organizational Affiliation: